Repression of the luteinizing hormone receptor gene promoter by cross talk among EAR3/COUP-TFI, Sp1/Sp3, and TFIIB
- PMID: 12972613
- PMCID: PMC193922
- DOI: 10.1128/MCB.23.19.6958-6972.2003
Repression of the luteinizing hormone receptor gene promoter by cross talk among EAR3/COUP-TFI, Sp1/Sp3, and TFIIB
Abstract
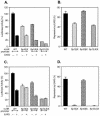
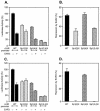
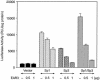
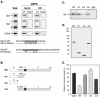
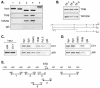
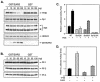
Transcription of luteinizing hormone receptor (LHR) gene is activated by Sp1/Sp3 at two Sp1 sites and is repressed by nuclear orphan receptors EAR2 and EAR3 through a direct-repeat (DR) motif. To elucidate the mechanism of the orphan receptor-mediated gene repression, we explored the functional connection between the orphan receptors and Sp1/Sp3 complex, and its impact on the basal transcription machinery. The Sp1(I) site was identified as critical for the repression since its mutation reduced the inhibition by EAR2 and abolished the inhibition by EAR3. Cotransfection analyses in SL2 cells showed that both Sp1 and Sp3 were required for this process since EAR3 displayed a complete Sp1/Sp3-dependent inhibitory effect. Functional cooperation between Sp1 and DR domains was further supported by mutual recruitment of EAR3 and Sp1/Sp3 bound to their cognate sites. Deletion of EAR3 N-terminal and DNA-binding domains that reduced its interaction with Sp1 impaired its inhibitory effect on human LHR (hLHR) gene transcription. Furthermore, we demonstrate interaction of TFIIB with Sp1/Sp3 at the Sp1(I) site besides its association with EAR3 and the TATA-less core promoter region. Such interaction relied on Sp1 site-bound Sp1/Sp3 complex and adaptor protein(s) present in the JAR nuclear extracts. We further demonstrated that EAR3 specifically decreased association of TFIIB to the Sp1(I) site without interfering on its interaction with the hLHR core promoter. The C-terminal region of EAR3, which did not participate in its interaction with Sp1, was required for its inhibitory function and may affect the association of TFIIB with Sp1. Moreover, perturbation of the association of TFIIB with Sp1 by EAR3 was reflected in the reduced recruitment of RNA polymerase II to the promoter. Overexpression of TFIIB counteracted the inhibitory effect of EAR3 and activated hLHR gene transcription in an Sp1 site-dependent manner. These findings therefore indicate that TFIIB is a key component in the regulatory control of EAR3 and Sp1/Sp3 on the initiation complex. Such cross talk among EAR3, TFIIB, and Sp1/Sp3 reveals repression of hLHR gene transcription by nuclear orphan receptors is achieved via perturbation of communication between Sp1/Sp3 at the Sp1-1 site and the basal transcription initiator complex.
Figures

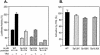
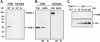
Similar articles
-
Dual mechanisms of regulation of transcription of luteinizing hormone receptor gene by nuclear orphan receptors and histone deacetylase complexes.J Steroid Biochem Mol Biol. 2003 Jun;85(2-5):401-14. doi: 10.1016/s0960-0760(03)00230-9. J Steroid Biochem Mol Biol. 2003. PMID: 12943729 Review.
-
EAR2 and EAR3/COUP-TFI regulate transcription of the rat LH receptor.Mol Endocrinol. 2001 Nov;15(11):1891-905. doi: 10.1210/mend.15.11.0720. Mol Endocrinol. 2001. PMID: 11682620
-
Nuclear orphan receptors regulate transcription of the gene for the human luteinizing hormone receptor.J Biol Chem. 2000 Jan 28;275(4):2763-70. doi: 10.1074/jbc.275.4.2763. J Biol Chem. 2000. PMID: 10644740
-
Silencing of transcription of the human luteinizing hormone receptor gene by histone deacetylase-mSin3A complex.J Biol Chem. 2002 Sep 6;277(36):33431-8. doi: 10.1074/jbc.M204417200. Epub 2002 Jun 28. J Biol Chem. 2002. PMID: 12091390
-
Participation of signaling pathways in the derepression of luteinizing hormone receptor transcription.Mol Cell Endocrinol. 2010 Jan 27;314(2):221-7. doi: 10.1016/j.mce.2009.05.005. Epub 2009 May 21. Mol Cell Endocrinol. 2010. PMID: 19464346 Free PMC article. Review.
Cited by
-
Nuclear receptor 4A1 (NR4A1) as a drug target for treating rhabdomyosarcoma (RMS).Oncotarget. 2016 May 24;7(21):31257-69. doi: 10.18632/oncotarget.9112. Oncotarget. 2016. PMID: 27144436 Free PMC article.
-
Coordinated changes in DNA methylation and histone modifications regulate silencing/derepression of luteinizing hormone receptor gene transcription.Mol Cell Biol. 2005 Sep;25(18):7929-39. doi: 10.1128/MCB.25.18.7929-7939.2005. Mol Cell Biol. 2005. PMID: 16135786 Free PMC article.
-
A role for Sp1 in transcriptional regulation of phosphatidylethanolamine N-methyltransferase in liver and 3T3-L1 adipocytes.J Biol Chem. 2010 Apr 16;285(16):11880-91. doi: 10.1074/jbc.M110.109843. Epub 2010 Feb 11. J Biol Chem. 2010. PMID: 20150657 Free PMC article.
-
Activation of Genes by Nuclear Receptor/Specificity Protein (Sp) Interactions in Cancer.Cancers (Basel). 2025 Jan 17;17(2):284. doi: 10.3390/cancers17020284. Cancers (Basel). 2025. PMID: 39858066 Free PMC article. Review.
-
Evidence of the neuron-restrictive silencer factor (NRSF) interaction with Sp3 and its synergic repression to the mu opioid receptor (MOR) gene.Nucleic Acids Res. 2006;34(22):6392-403. doi: 10.1093/nar/gkl724. Epub 2006 Nov 27. Nucleic Acids Res. 2006. PMID: 17130167 Free PMC article.
References
-
- Bailey, P. J., D. H. Dowhan, K. Franke, L. J. Burke, M. Downes, and G. E. Muscat. 1997. Transcriptional repression by COUP-TF II is dependent on the C-terminal domain and involves the N-CoR variant, RIP13delta1. J. Steroid Biochem. Mol. Biol. 63:165-174. - PubMed
-
- Black, A. R., J. D. Black, and J. Azizkhan-Clifford. 2001. Sp1 and Kruppel-like factor family of transcription factors in cell growth regulation and cancer. J. Cell Physiol. 188:143-160. - PubMed
-
- Campbell, A. 2002. DNA binding and bending to initiate packaging of phage lambda DNA. Mol. Cell 9:928-929. - PubMed
-
- Catt, K. J., and M. L. Dufau. 1991. Gonadotropic hormones: biosynthesis, secretion, receptors and action, p. 105-155. In S. S. C. Yen and R. B. Jaffe (ed.), Reproductive endocrinology. The W. B. Saunders Co., Philadelphia, Pa.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous
