Unique Sm core structure of U7 snRNPs: assembly by a specialized SMN complex and the role of a new component, Lsm11, in histone RNA processing
- PMID: 12975319
- PMCID: PMC196468
- DOI: 10.1101/gad.274403
Unique Sm core structure of U7 snRNPs: assembly by a specialized SMN complex and the role of a new component, Lsm11, in histone RNA processing
Abstract
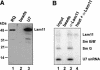
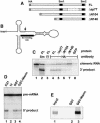
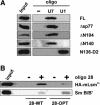
A set of seven Sm proteins assemble on the Sm-binding site of spliceosomal U snRNAs to form the ring-shaped Sm core. The U7 snRNP involved in histone RNA 3' processing contains a structurally similar but biochemically unique Sm core in which two of these proteins, Sm D1 and D2, are replaced by Lsm10 and by another as yet unknown component. Here we characterize this factor, termed Lsm11, as a novel Sm-like protein with apparently two distinct functions. In vitro studies suggest that its long N-terminal part mediates an important step in histone mRNA 3'-end cleavage, most likely by recruiting a zinc finger protein previously identified as a processing factor. In contrast, the C-terminal part, which comprises two Sm motifs interrupted by an unusually long spacer, is sufficient to assemble with U7, but not U1, snRNA. Assembly of this U7-specific Sm core depends on the noncanonical Sm-binding site of U7 snRNA. Moreover, it is facilitated by a specialized SMN complex that contains Lsm10 and Lsm11 but lacks Sm D1/D2. Thus, the U7-specific Lsm11 protein not only specifies the assembly of the U7 Sm core but also fulfills an important role in U7 snRNP-mediated histone mRNA processing.
Figures

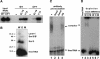

References
-
- Brahms H., Raymackers, J., Union, A., de Keyser, F., Meheus, L., and Lührmann, R. 2000. The C-terminal RG dipeptide repeats of the spliceosomal Sm proteins D1 and D3 contain symmetrical dimethylarginines, which form a major B-cell epitope for anti-Sm autoantibodies. J. Biol. Chem. 275: 17122-17129. - PubMed
-
- Chittum H.S., Lane, W.S., Carlson, B.A., Roller, P.P., Lung, F.D., Lee, B.J., and Hatfield, D.L. 1998. Rabbit β-globin is extended beyond its UGA stop codon by multiple suppressions and translational reading gaps. Biochemistry 37: 10866-10870. - PubMed
-
- Dominski Z. and Marzluff, W.F. 1999. Formation of the 3′ end of histone mRNA. Gene 239: 1-14. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
