Reversible and fast association equilibria of a molecular chaperone, gp57A, of bacteriophage T4
- PMID: 14507723
- PMCID: PMC1303484
- DOI: 10.1016/s0006-3495(03)74683-9
Reversible and fast association equilibria of a molecular chaperone, gp57A, of bacteriophage T4
Abstract
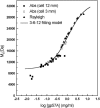
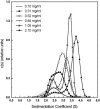
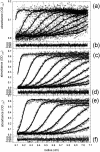
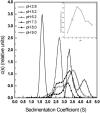
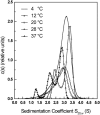
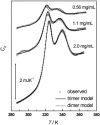
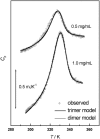
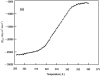
The association of a molecular chaperone, gp57A, of bacteriophage T4, which facilitates formation of the long and short tail fibers, was investigated by analytical ultracentrifugation, differential scanning microcalorimetry, and stopped-flow circular dichroism (CD) to establish the association scheme of the protein. Gp57A is an oligomeric alpha-helix protein with 79 amino acids. Analysis of the sedimentation velocity data by direct boundary modeling with Lamm equation solutions together with a more detailed boundary analysis incorporating association schemes led us to conclude that at least three oligomeric species of gp57A are in reversible and fast association equilibria and that a 3(mer)-6(mer)-12(mer) model described the data best. On the other hand, differential scanning microcalorimetry revealed a highly reversible two-step transition of dissociation/denaturation, both of which accompanied decrease in CD at 222 nm. The melting curve analysis revealed that it is consistent with a 6(mer)-3(mer)-1(mer) model. The refolding/association kinetics of gp57A measured by stopped-flow CD was consistent with the interpretation that the bimolecular reaction from trimer to hexamer was preceded by a fast alpha-helix formation in the dead-time. Trimer or hexamer is likely the functional oligomeric state of gp57A.
Figures


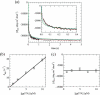
References
-
- Arai, M., and K. Kuwajima. 1996. Rapid formation of a molten globule intermediate in refolding of α-lactalbumin. Fold. Des. 1:275–287. - PubMed
-
- Bishop, J. R., M. P. Conley, and W. B. Wood. 1974. Assembly and attachment of bacteriophage T4 tail fibers. J. Supramol. Struct. 2:196–201. - PubMed
-
- Burda, M. R., and S. Miller. 1999. Folding of coliphage T4 short tail fiber in vitro. Analysing the role of a bacteriophage-encoded chaperone. Eur. J. Biochem. 265:771–778. - PubMed
-
- Cox, D. J. 1969. Computer simulation of sedimentation in the ultracentrifuge. IV. Velocity sedimentation of self-associating solutes. Arch. Biochem. Biophys. 129:106–123. - PubMed
-
- Dam, J., and P. Schuck. 2003. Calculating sedimentation coefficient distributions by direct modeling of sedimentation velocity data. Methods Enzymol. In press. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

