Patterns of community change among ammonia oxidizers in meadow soils upon long-term incubation at different temperatures
- PMID: 14532075
- PMCID: PMC201211
- DOI: 10.1128/AEM.69.10.6152-6164.2003
Patterns of community change among ammonia oxidizers in meadow soils upon long-term incubation at different temperatures
Abstract
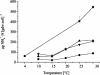
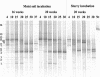
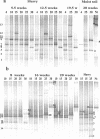
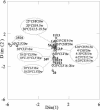
The effect of temperature on the community structure of ammonia-oxidizing bacteria was investigated in three different meadow soils. Two of the soils (OMS and GMS) were acidic (pH 5.0 to 5.8) and from sites in Germany with low annual mean temperature (about 10 degrees C), while KMS soil was slightly alkaline (pH 7.9) and from a site in Israel with a high annual mean temperature (about 22 degrees C). The soils were fertilized and incubated for up to 20 weeks in a moist state and as a buffered (pH 7) slurry amended with urea at different incubation temperatures (4 to 37 degrees C). OMS soil was also incubated with less fertilizer than the other soils. The community structure of ammonia oxidizers was analyzed before and after incubation by denaturing gradient gel electrophoresis (DGGE) of the amoA gene, which codes for the alpha subunit of ammonia monooxygenase. All amoA gene sequences found belonged to the genus Nitrosospira. The analysis showed community change due to temperature both in moist soil and in the soil slurry. Two patterns of community change were observed. One pattern was a change between the different Nitrosospira clusters, which was observed in moist soil and slurry incubations of GMS and OMS. Nitrosospira AmoA cluster 1 was mainly detected below 30 degrees C, while Nitrosospira cluster 4 was predominant at 25 degrees C. Nitrosospira clusters 3a, 3b, and 9 dominated at 30 degrees C. The second pattern, observed in KMS, showed a community shift predominantly within a single Nitrosospira cluster. The sequences of the individual DGGE bands that exhibited different trends with temperature belonged almost exclusively to Nitrosospira cluster 3a. We conclude that ammonia oxidizer populations are influenced by temperature. In addition, we confirmed previous observations that N fertilizer also influences the community structure of ammonia oxidizers. Thus, Nitrosospira cluster 1 was absent in OMS soil treated with less fertilizer, while Nitrosospira cluster 9 was only found in the sample given less fertilizer.
Figures






References
-
- Aakra, A., J. B. Utaker, and I. F. Nes. 2001. Comparative phylogeny of the ammonia monooxygenase subunit A and 16S rRNA genes of ammonia-oxidizing bacteria. FEMS Microbiol. Lett. 205:237-242. - PubMed
-
- Aakra, A., J. B. Utaker, I. F. Nes, and L. R. Bakken. 1999. An evaluated improvement of the extinction dilution method for isolation of ammonia-oxidizing bacteria. J. Microbiol. Methods 39:23-31. - PubMed
-
- Avrahami, S., W. Liesack, and R. Conrad. 2003. Effects of temperature and fertilizer on activity and community structure of soil ammonia oxidizers. Environ. Microbiol. 5:691-705. - PubMed
-
- Bak, F., G. Scheff, and K. H. Jansen. 1991. A rapid and sensitive ion chromatographic technique for the determination of sulfate and sulfate reduction rates in freshwater lake sediments. FEMS Microbiol. Ecol. 85:23-30.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

