Coronaviruses as vectors: position dependence of foreign gene expression
- PMID: 14557617
- PMCID: PMC229330
- DOI: 10.1128/jvi.77.21.11312-11323.2003
Coronaviruses as vectors: position dependence of foreign gene expression
Abstract
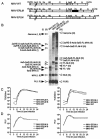
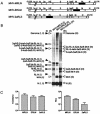
Coronaviruses are the enveloped, positive-stranded RNA viruses with the largest RNA genomes known. Several features make these viruses attractive as vaccine and therapeutic vectors: (i) deletion of their nonessential genes is strongly attenuating; (ii) the genetic space thus created allows insertion of foreign information; and (iii) their tropism can be modified by manipulation of the viral spike. We studied here their ability to serve as expression vectors by inserting two different foreign genes and evaluating systematically the genomic position dependence of their expression, using a murine coronavirus as a model. Renilla and firefly luciferase expression cassettes, each provided with viral transcription regulatory sequences (TRSs), were inserted at several genomic positions, both independently in different viruses and combined within one viral genome. Recombinant viruses were generated by using a convenient method based on targeted recombination and host cell switching. In all cases high expression levels of the foreign genes were observed without severe effects on viral replication in vitro. The expression of the inserted gene appeared to be dependent on its genomic position, as well as on the identity of the gene. Expression levels increased when the luciferase gene was inserted closer to the 3' end of the genome. The foreign gene insertions generally reduced the expression of upstream viral genes. The results are consistent with coronavirus transcription models in which the transcription from upstream TRSs is attenuated by downstream TRSs. Altogether, our observations clearly demonstrate the potential of coronaviruses as (multivalent) expression vectors.
Figures



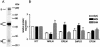


References
-
- Bredenbeek, P. J., C. J. Pachuk, A. F. Noten, J. Charite, W. Luytjes, S. R. Weiss, and W. J. Spaan. 1990. The primary structure and expression of the second open reading frame of the polymerase gene of the coronavirus MHV-A59; a highly conserved polymerase is expressed by an efficient ribosomal frameshifting mechanism. Nucleic Acids Res. 18:1825-1832. - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

