Molecular basis of passive stress relaxation in human soleus fibers: assessment of the role of immunoglobulin-like domain unfolding
- PMID: 14581214
- PMCID: PMC1303590
- DOI: 10.1016/S0006-3495(03)74732-8
Molecular basis of passive stress relaxation in human soleus fibers: assessment of the role of immunoglobulin-like domain unfolding
Abstract
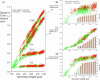
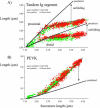
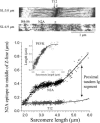
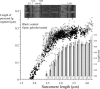
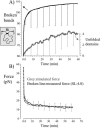
Titin (also known as connectin) is the main determinant of physiological levels of passive muscle force. This force is generated by the extensible I-band region of the molecule, which is constructed of the PEVK domain and tandem-immunoglobulin segments comprising serially linked immunoglobulin (Ig)-like domains. It is unresolved whether under physiological conditions Ig domains remain folded and act as "spacers" that set the sarcomere length at which the PEVK extends or whether they contribute to titin's extensibility by unfolding. Here we focused on whether Ig unfolding plays a prominent role in stress relaxation (decay of force at constant length after stretch) using mechanical and immunolabeling studies on relaxed human soleus muscle fibers and Monte Carlo simulations. Simulation experiments using Ig-domain unfolding parameters obtained in earlier single-molecule atomic force microscopy experiments recover the phenomenology of stress relaxation and predict large-scale unfolding in titin during an extended period (> approximately 20 min) of relaxation. By contrast, immunolabeling experiments failed to demonstrate large-scale unfolding. Thus, under physiological conditions in relaxed human soleus fibers, Ig domains are more stable than predicted by atomic force microscopy experiments. Ig-domain unfolding did not become more pronounced after gelsolin treatment, suggesting that the thin filament is unlikely to significantly contribute to the mechanical stability of the domains. We conclude that in human soleus fibers, Ig unfolding cannot solely explain stress relaxation.
Figures





Similar articles
-
Titin extensibility in situ: entropic elasticity of permanently folded and permanently unfolded molecular segments.J Cell Biol. 1998 Feb 23;140(4):853-9. doi: 10.1083/jcb.140.4.853. J Cell Biol. 1998. PMID: 9472037 Free PMC article.
-
Unfolding of titin domains explains the viscoelastic behavior of skeletal myofibrils.Biophys J. 2001 Mar;80(3):1442-51. doi: 10.1016/S0006-3495(01)76116-4. Biophys J. 2001. PMID: 11222304 Free PMC article.
-
PEVK extension of human soleus muscle titin revealed by immunolabeling with the anti-titin antibody 9D10.J Struct Biol. 1998;122(1-2):188-96. doi: 10.1006/jsbi.1998.3984. J Struct Biol. 1998. PMID: 9724620
-
Titin elasticity in the context of the sarcomere: force and extensibility measurements on single myofibrils.Adv Exp Med Biol. 2000;481:179-202; discussion 203-6. doi: 10.1007/978-1-4615-4267-4_11. Adv Exp Med Biol. 2000. PMID: 10987073 Review.
-
Stretching molecular springs: elasticity of titin filaments in vertebrate striated muscle.Histol Histopathol. 2000 Jul;15(3):799-811. doi: 10.14670/HH-15.799. Histol Histopathol. 2000. PMID: 10963124 Review.
Cited by
-
Titin-actin interaction: PEVK-actin-based viscosity in a large animal.J Biomed Biotechnol. 2011;2011:310791. doi: 10.1155/2011/310791. Epub 2011 Nov 15. J Biomed Biotechnol. 2011. PMID: 22162634 Free PMC article.
-
Passive force and viscoelastic properties of single fibers in human aging muscles.Eur J Appl Physiol. 2019 Oct;119(10):2339-2348. doi: 10.1007/s00421-019-04221-7. Epub 2019 Aug 29. Eur J Appl Physiol. 2019. PMID: 31468173
-
Pulling single molecules of titin by AFM--recent advances and physiological implications.Pflugers Arch. 2008 Apr;456(1):101-15. doi: 10.1007/s00424-007-0389-x. Epub 2007 Dec 6. Pflugers Arch. 2008. PMID: 18058125 Review.
-
Can the passive elasticity of muscle be explained directly from the mechanics of individual titin molecules?J Muscle Res Cell Motil. 2005;26(6-8):285-9. doi: 10.1007/s10974-005-9034-5. J Muscle Res Cell Motil. 2005. PMID: 16465473
-
Activation and stretch-induced passive force enhancement--are you pulling my chain? Focus on "Regulation of muscle force in the absence of actin-myosin-based cross-bridge interaction".Am J Physiol Cell Physiol. 2010 Jul;299(1):C11-3. doi: 10.1152/ajpcell.00147.2010. Epub 2010 May 5. Am J Physiol Cell Physiol. 2010. PMID: 20445175 Free PMC article. No abstract available.
References
-
- Bang, M. L., T. Centner, F. Fornoff, A. J. Geach, M. Gotthardt, M. McNabb, C. C. Witt, D. Labeit, C. C. Gregorio, H. Granzier, and S. Labeit. 2001. The complete gene sequence of titin, expression of an unusual approximately 700-kDa titin isoform, and its interaction with obscurin identify a novel Z-line to I-band linking system. Circ. Res. 89:1065–1072. - PubMed
-
- Bustamante, C., J. F. Marko, E. D. Siggia, and S. Smith. 1994. Entropic elasticity of lambda-phage DNA. Science. 265:1599–1600. - PubMed
-
- Carrion-Vazquez, M., A. F. Oberhauser, T. E. Fisher, P. E. Marszalek, H. Li, and J. M. Fernandez. 2000. Mechanical design of proteins studied by single-molecule force spectroscopy and protein engineering. Prog. Biophys. Mol. Biol. 74:63–91. - PubMed
-
- Cazorla, O., A. Freiburg, M. Helmes, T. Centner, M. McNabb, Y. Wu, K. Trombitas, S. Labeit, and H. Granzier. 2000. Differential expression of cardiac titin isoforms and modulation of cellular stiffness. Circ. Res. 86:59–67. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

