Identification of 13 novel human modification guide RNAs
- PMID: 14602913
- PMCID: PMC275545
- DOI: 10.1093/nar/gkg849
Identification of 13 novel human modification guide RNAs
Abstract
Members of the two expanding RNA subclasses termed C/D and H/ACA RNAs guide the 2'-O-methylations and pseudouridylations, respectively, of rRNA and spliceosomal RNAs (snRNAs). Here, we report on the identification of 13 novel human intron-encoded small RNAs (U94-U106) belonging to the two subclasses of modification guides. Seven of them are predicted to direct 2'-O-methylations in rRNA or snRNAs, while the remainder represent novel orphan RNA modification guides. From these, U100, which is exclusively detected in Cajal bodies (CBs), is predicted to direct modification of a U6 snRNA uridine, U(9), which to date has not been found to be pseudouridylated. Hence, within CBs, U100 might function in the folding pathway or other aspects of U6 snRNA metabolism rather than acting as a pseudouridylation guide. U106 C/D snoRNA might also possess an RNA chaperone activity only since its two conserved antisense elements match two rRNA sequences devoid of methylated nucleotides and located remarkably close to each other within the 18S rRNA secondary structure. Finally, we have identified a retrogene for U99 snoRNA located within an intron of the Siat5 gene, supporting the notion that retro-transposition events might have played a substantial role in the mobility and diversification of snoRNA genes during evolution.
Figures




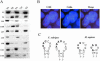

References
-
- Eddy S.R. (2001) Non-coding RNA genes and the modern RNA world. Nature Rev. Genet., 2, 919–929. - PubMed
-
- Mattick J.S. and Gagen,M.J. (2001) The evolution of controlled multitasked gene networks: the role of introns and other noncoding RNAs in the development of complex organisms. Mol. Biol. Evol., 18, 1611–1630. - PubMed
-
- Wassarman K.M. (2002) Small RNAs in bacteria: diverse regulators of gene expression in response to environmental changes. Cell, 109, 141–144. - PubMed
-
- Kiss-Laszlo Z., Henry,Y., Bachellerie,J.P., Caizergues-Ferrer,M. and Kiss,T. (1996) Site-specific ribose methylation of preribosomal RNA: a novel function for small nucleolar RNAs. Cell, 85, 1077–1088. - PubMed
-
- Ganot P., Caizergues-Ferrer,M. and Kiss,T. (1997) The family of box ACA small nucleolar RNAs is defined by an evolutionarily conserved secondary structure and ubiquitous sequence elements essential for RNA accumulation. Genes Dev., 11, 941–956. - PubMed

