Dimerization controls the lipid raft partitioning of uPAR/CD87 and regulates its biological functions
- PMID: 14609946
- PMCID: PMC275445
- DOI: 10.1093/emboj/cdg588
Dimerization controls the lipid raft partitioning of uPAR/CD87 and regulates its biological functions
Abstract
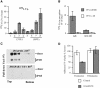
The urokinase-type plasminogen activator receptor (uPAR/CD87) is a glycosylphosphatidylinositol-anchored membrane protein with multiple functions in extracellular proteolysis, cell adhesion, cell migration and proliferation. We now report that cell surface uPAR dimerizes and that dimeric uPAR partitions preferentially to detergent-resistant lipid rafts. Dimerization of uPAR did not require raft partitioning as the lowering of membrane cholesterol failed to reduce dimerization and as a transmembrane uPAR chimera, which does not partition to lipid rafts, also dimerized efficiently. While uPA bound to uPAR independently of its membrane localization and dimerization status, uPA-induced uPAR cleavage was strongly accelerated in lipid rafts. In contrast to uPA, the binding of Vn occurred preferentially to raft- associated dimeric uPAR and was completely blocked by cholesterol depletion.
Figures






Similar articles
-
Urokinase-receptor-mediated phenotypic changes in vascular smooth muscle cells require the involvement of membrane rafts.Biochem J. 2009 Oct 12;423(3):343-51. doi: 10.1042/BJ20090447. Biochem J. 2009. PMID: 19691446
-
Lipid raft compartmentalization of urokinase receptor signaling in human neutrophils.Am J Respir Cell Mol Biol. 2004 Feb;30(2):233-41. doi: 10.1165/rcmb.2003-0079OC. Epub 2003 Aug 21. Am J Respir Cell Mol Biol. 2004. PMID: 12933356
-
Cyclo19,31[D-Cys19]-uPA19-31 is a potent competitive antagonist of the interaction of urokinase-type plasminogen activator with its receptor (CD87).Biol Chem. 2001 Aug;382(8):1197-205. doi: 10.1515/BC.2001.150. Biol Chem. 2001. PMID: 11592401
-
Structure, function and expression on blood and bone marrow cells of the urokinase-type plasminogen activator receptor, uPAR.Stem Cells. 1997;15(6):398-408. doi: 10.1002/stem.150398. Stem Cells. 1997. PMID: 9402652 Review.
-
The structure and function of the urokinase receptor, a membrane protein governing plasminogen activation on the cell surface.Biol Chem Hoppe Seyler. 1995 May;376(5):269-79. Biol Chem Hoppe Seyler. 1995. PMID: 7662169 Review.
Cited by
-
Bilayer asymmetry influences integrin sequestering in raft-mimicking lipid mixtures.Biophys J. 2013 May 21;104(10):2212-21. doi: 10.1016/j.bpj.2013.04.020. Biophys J. 2013. PMID: 23708361 Free PMC article.
-
The cross-talk between the urokinase receptor and fMLP receptors regulates the activity of the CXCR4 chemokine receptor.Cell Mol Life Sci. 2011 Jul;68(14):2453-67. doi: 10.1007/s00018-010-0564-7. Epub 2010 Oct 24. Cell Mol Life Sci. 2011. PMID: 20972812 Free PMC article.
-
Clathrin and LRP-1-independent constitutive endocytosis and recycling of uPAR.PLoS One. 2008;3(11):e3730. doi: 10.1371/journal.pone.0003730. Epub 2008 Nov 14. PLoS One. 2008. PMID: 19008962 Free PMC article.
-
Differential uPAR recruitment in caveolar-lipid rafts by GM1 and GM3 gangliosides regulates endothelial progenitor cells angiogenesis.J Cell Mol Med. 2015 Jan;19(1):113-23. doi: 10.1111/jcmm.12410. Epub 2014 Oct 14. J Cell Mol Med. 2015. PMID: 25313007 Free PMC article.
-
New Pieces in the Puzzle of uPAR Role in Cell Migration Mechanisms.Cells. 2020 Nov 24;9(12):2531. doi: 10.3390/cells9122531. Cells. 2020. PMID: 33255171 Free PMC article.
References
-
- Andolfo A., English,W.R., Resnati,M., Murphy,G., Blasi,F. and Sidenius,N. (2002) Metalloproteases cleave the urokinase-type plasminogen activator receptor in the D1–D2 linker region and expose epitopes not present in the intact soluble receptor. Thromb. Haemost., 88, 298–306. - PubMed
-
- Behrendt N., Ploug,M., Patthy,L., Houen,G., Blasi,F. and Danø,K. (1991) The ligand-binding domain of the cell surface receptor for urokinase-type plasminogen activator. J. Biol. Chem., 266, 7842–7847. - PubMed
-
- Blasi F. and Carmeliet,P. (2002) uPAR: a versatile signalling orchestrator. Nat. Rev. Mol. Cell Biol., 3, 932–943. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

