Site-1 protease is required for cartilage development in zebrafish
- PMID: 14612568
- PMCID: PMC283539
- DOI: 10.1073/pnas.2331794100
Site-1 protease is required for cartilage development in zebrafish
Abstract
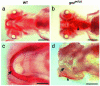
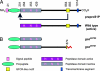
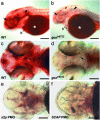
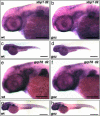
gonzo (goz) is a zebrafish mutant with defects in cartilage formation. The goz phenotype comprises cartilage matrix defects and irregular chondrocyte morphology. Expression of endoderm, mesoderm, and cartilage marker genes is, however, normal, indicating a defect in chondrocyte morphogenesis. The mutated gene responsible for the goz phenotype, identified by positional cloning and confirmed by phosphomorpholino knockdown, encodes zebrafish site-1 protease (s1p). S1P has been shown to process and activate sterol regulatory element-binding proteins (SREBPs), which regulate expression of key enzymes of lipid biosynthesis or transport. This finding is consistent with the abnormal distribution of lipids in goz embryos. Knockdown of site-2 protease, which is also involved in activation of SREBPs, results in similar lipid and cartilage phenotypes as S1P knockdown. However, knockdown of SREBP cleavage-activating protein, which forms a complex with SREBP and is essential for S1P cleavage, results only in lipid phenotypes, whereas cartilage appears normal. This indicates that the cartilage phenoptypes of goz are caused independently of the lipid defects.
Figures





Similar articles
-
Proteolytic activation of SREBPs during adipocyte differentiation.Biochem Biophys Res Commun. 2001 May 25;283(5):1157-61. doi: 10.1006/bbrc.2001.4915. Biochem Biophys Res Commun. 2001. PMID: 11355894
-
Decreased lipid synthesis in livers of mice with disrupted Site-1 protease gene.Proc Natl Acad Sci U S A. 2001 Nov 20;98(24):13607-12. doi: 10.1073/pnas.201524598. Proc Natl Acad Sci U S A. 2001. PMID: 11717426 Free PMC article.
-
ER stress induces cleavage of membrane-bound ATF6 by the same proteases that process SREBPs.Mol Cell. 2000 Dec;6(6):1355-64. doi: 10.1016/s1097-2765(00)00133-7. Mol Cell. 2000. PMID: 11163209
-
[Site-1 protease: a subtilisin-like serine protease that cleaves SREBPs for controlling the lipid biosynthesis in animal cell].Nihon Rinsho. 2001 Feb;59 Suppl 2:277-82. Nihon Rinsho. 2001. PMID: 11351588 Review. Japanese. No abstract available.
-
A proteolytic pathway that controls the cholesterol content of membranes, cells, and blood.Proc Natl Acad Sci U S A. 1999 Sep 28;96(20):11041-8. doi: 10.1073/pnas.96.20.11041. Proc Natl Acad Sci U S A. 1999. PMID: 10500120 Free PMC article. Review.
Cited by
-
Exome Sequencing Reveals Biallelic Mutations in MBTPS1 Gene in a Girl with a Very Rare Skeletal Dysplasia.Diagnostics (Basel). 2024 Jan 31;14(3):313. doi: 10.3390/diagnostics14030313. Diagnostics (Basel). 2024. PMID: 38337829 Free PMC article.
-
Gut-like ectodermal tissue in a sea anemone challenges germ layer homology.Nat Ecol Evol. 2017 Oct;1(10):1535-1542. doi: 10.1038/s41559-017-0285-5. Epub 2017 Sep 11. Nat Ecol Evol. 2017. PMID: 29185520 Free PMC article.
-
Genetically engineered zebrafish as models of skeletal development and regeneration.Bone. 2023 Feb;167:116611. doi: 10.1016/j.bone.2022.116611. Epub 2022 Nov 14. Bone. 2023. PMID: 36395960 Free PMC article. Review.
-
2,4-Dinitrotoluene (DNT) Perturbs Yolk Absorption, Liver Development and Lipid Metabolism/Oxygen Transport Gene Expression in Zebrafish Embryos and Larvae.Int J Mol Sci. 2019 Jul 25;20(15):3632. doi: 10.3390/ijms20153632. Int J Mol Sci. 2019. PMID: 31349543 Free PMC article.
-
Site-1 protease regulates skeletal stem cell population and osteogenic differentiation in mice.Biol Open. 2018 Feb 22;7(2):bio032094. doi: 10.1242/bio.032094. Biol Open. 2018. PMID: 29437042 Free PMC article.
References
-
- Olson, E. N., Brown, D., Burgess, R. & Cserjesi, P. (1996) Ann. N. Y. Acad. Sci. 785, 108–118. - PubMed
-
- Zhang, Z., Song, Y., Zhao, X., Zhang, X., Fermin, C. & Chen, Y. (2002) Development (Cambridge, U.K.) 129, 4135–4146. - PubMed
-
- ten Berge, D., Brouwer, A., Korving, J., Reijnen, M. J., van Raaij, E. J., Verbeek, F., Gaffield, W. & Meijlink, F. (2001) Development (Cambridge, U.K.) 128, 2929–2938. - PubMed
-
- Liu, W., Li, G., Chien, J. S., Raft, S., Zhang, H., Chiang, C. & Frenz, D. A. (2002) Dev. Biol. 248, 240–250. - PubMed
-
- Hu, D. & Helms, J. A. (1999) Development (Cambridge, U.K.) 126, 4873–4884. - PubMed
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

