Backtracking by single RNA polymerase molecules observed at near-base-pair resolution
- PMID: 14634670
- PMCID: PMC1483218
- DOI: 10.1038/nature02191
Backtracking by single RNA polymerase molecules observed at near-base-pair resolution
Abstract
Escherichia coli RNA polymerase (RNAP) synthesizes RNA with remarkable fidelity in vivo. Its low error rate may be achieved by means of a 'proofreading' mechanism comprised of two sequential events. The first event (backtracking) involves a transcriptionally upstream motion of RNAP through several base pairs, which carries the 3' end of the nascent RNA transcript away from the enzyme active site. The second event (endonucleolytic cleavage) occurs after a variable delay and results in the scission and release of the most recently incorporated ribonucleotides, freeing up the active site. Here, by combining ultrastable optical trapping apparatus with a novel two-bead assay to monitor transcriptional elongation with near-base-pair precision, we observed backtracking and recovery by single molecules of RNAP. Backtracking events ( approximately 5 bp) occurred infrequently at locations throughout the DNA template and were associated with pauses lasting 20 s to >30 min. Inosine triphosphate increased the frequency of backtracking pauses, whereas the accessory proteins GreA and GreB, which stimulate the cleavage of nascent RNA, decreased the duration of such pauses.
Figures
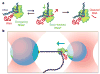
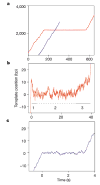

References
-
- Erie DA, Yager TD, von Hippel PH. The single-nucleotide addition cycle in transcription: a biophysical and biochemical perspective. Annu Rev Biophys Biomol Struct. 1992;21:379–415. - PubMed
-
- Thomas MJ, Platas AA, Hawley DK. Transcriptional fidelity and proofreading by RNA polymerase II. Cell. 1998;93:627–637. - PubMed
-
- Erie DA, Hajiseyedjavadi O, Young MC, von Hippel PH. Multiple RNA polymerase conformations and GreA: control of the fidelity of transcription. Science. 1993;262:867–873. - PubMed
-
- Kunkel TA, Bebenek K. DNA replication fidelity. Annu Rev Biochem. 2000;69:497–529. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

