Structure and function of AMPA receptors
- PMID: 14645452
- PMCID: PMC1664757
- DOI: 10.1113/jphysiol.2003.054320
Structure and function of AMPA receptors
Abstract
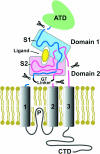
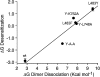
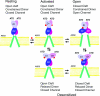
The application of X-ray diffraction has allowed the structure of the ligand-binding core of AMPA receptors to be determined. Here I review the insights that this has given into the molecular mechanisms of activation and desensitization of these receptors.
Figures
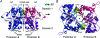

Similar articles
-
Structural aspects of AMPA receptor activation, desensitization and deactivation.Curr Opin Neurobiol. 2007 Jun;17(3):281-8. doi: 10.1016/j.conb.2007.03.014. Epub 2007 Apr 6. Curr Opin Neurobiol. 2007. PMID: 17419047 Review.
-
The inner workings of the AMPA receptors.Curr Opin Drug Discov Devel. 2002 Sep;5(5):741-8. Curr Opin Drug Discov Devel. 2002. PMID: 12630294 Review.
-
Mechanisms of channel gating of the ligand-gated ion channel superfamily inferred from protein structure.Exp Physiol. 2004 Mar;89(2):145-53. doi: 10.1113/expphysiol.2003.026815. Epub 2004 Feb 17. Exp Physiol. 2004. PMID: 15123543 Review.
-
Mechanism of inhibition of the GluR2 AMPA receptor channel opening by 2,3-benzodiazepine derivatives.Biochemistry. 2008 Jan 22;47(3):1061-9. doi: 10.1021/bi700782x. Epub 2007 Dec 28. Biochemistry. 2008. PMID: 18161947
-
Teaching resources. Ligand-gated ion channels.Sci STKE. 2005 Apr 19;2005(280):tr12. doi: 10.1126/stke.2802005tr12. Sci STKE. 2005. PMID: 15840839
Cited by
-
Structure of the Arabidopsis Glutamate Receptor-like Channel GLR3.2 Ligand-Binding Domain.Structure. 2021 Feb 4;29(2):161-169.e4. doi: 10.1016/j.str.2020.09.006. Epub 2020 Oct 6. Structure. 2021. PMID: 33027636 Free PMC article.
-
Role of the Ion Channel Extracellular Collar in AMPA Receptor Gating.Sci Rep. 2017 Apr 21;7(1):1050. doi: 10.1038/s41598-017-01146-z. Sci Rep. 2017. PMID: 28432359 Free PMC article.
-
LRET investigations of conformational changes in the ligand binding domain of a functional AMPA receptor.Biochemistry. 2008 Sep 23;47(38):10027-32. doi: 10.1021/bi800690b. Epub 2008 Aug 30. Biochemistry. 2008. PMID: 18759455 Free PMC article.
-
Role of conformational dynamics in α-amino-3-hydroxy-5-methylisoxazole-4-propionic acid (AMPA) receptor partial agonism.J Biol Chem. 2012 Dec 21;287(52):43557-64. doi: 10.1074/jbc.M112.371815. Epub 2012 Oct 31. J Biol Chem. 2012. PMID: 23115239 Free PMC article.
-
Cyclothiazide-induced persistent increase in respiratory-related activity in vitro.J Physiol. 2012 Oct 1;590(19):4897-915. doi: 10.1113/jphysiol.2012.232421. Epub 2012 Jul 2. J Physiol. 2012. PMID: 22753547 Free PMC article.
References
-
- Armstrong N, Gouaux E. Mechanisms for activation and antagonism of an AMPA-sensitive glutamate receptor: Crystal structures of the GluR2 ligand binding core. Neuron. 2000;28:165–181. - PubMed
-
- Armstrong N, Sun Y, Chen G-Q, Gouaux E. Structure of a glutamate receptor ligand binding core in complex with kainate. Nature. 1998;395:913–917. - PubMed
-
- Dingledine R, Borges K, Bowie D, Traynelis SF. The glutamate receptor ion channels. Pharmacol Rev. 1999;51:7–61. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

