Tumor classification and marker gene prediction by feature selection and fuzzy c-means clustering using microarray data
- PMID: 14651757
- PMCID: PMC302113
- DOI: 10.1186/1471-2105-4-60
Tumor classification and marker gene prediction by feature selection and fuzzy c-means clustering using microarray data
Abstract
Background: Using DNA microarrays, we have developed two novel models for tumor classification and target gene prediction. First, gene expression profiles are summarized by optimally selected Self-Organizing Maps (SOMs), followed by tumor sample classification by Fuzzy C-means clustering. Then, the prediction of marker genes is accomplished by either manual feature selection (visualizing the weighted/mean SOM component plane) or automatic feature selection (by pair-wise Fisher's linear discriminant).
Results: The proposed models were tested on four published datasets: (1) Leukemia (2) Colon cancer (3) Brain tumors and (4) NCI cancer cell lines. The models gave class prediction with markedly reduced error rates compared to other class prediction approaches, and the importance of feature selection on microarray data analysis was also emphasized.
Conclusions: Our models identify marker genes with predictive potential, often better than other available methods in the literature. The models are potentially useful for medical diagnostics and may reveal some insights into cancer classification. Additionally, we illustrated two limitations in tumor classification from microarray data related to the biology underlying the data, in terms of (1) the class size of data, and (2) the internal structure of classes. These limitations are not specific for the classification models used.
Figures

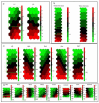



Similar articles
-
An unsupervised hierarchical dynamic self-organizing approach to cancer class discovery and marker gene identification in microarray data.Bioinformatics. 2003 Nov 1;19(16):2131-40. doi: 10.1093/bioinformatics/btg296. Bioinformatics. 2003. PMID: 14594719
-
Tumor classification by partial least squares using microarray gene expression data.Bioinformatics. 2002 Jan;18(1):39-50. doi: 10.1093/bioinformatics/18.1.39. Bioinformatics. 2002. PMID: 11836210
-
SamCluster: an integrated scheme for automatic discovery of sample classes using gene expression profile.Bioinformatics. 2003 May 1;19(7):811-7. doi: 10.1093/bioinformatics/btg095. Bioinformatics. 2003. PMID: 12724290
-
DNA microarrays in clinical cancer research.Curr Mol Med. 2005 Feb;5(1):111-20. doi: 10.2174/1566524053152834. Curr Mol Med. 2005. PMID: 15720274 Review.
-
Pathological bases for a robust application of cancer molecular classification.Int J Mol Sci. 2015 Apr 17;16(4):8655-75. doi: 10.3390/ijms16048655. Int J Mol Sci. 2015. PMID: 25898411 Free PMC article. Review.
Cited by
-
Instance-based concept learning from multiclass DNA microarray data.BMC Bioinformatics. 2006 Feb 16;7:73. doi: 10.1186/1471-2105-7-73. BMC Bioinformatics. 2006. PMID: 16483361 Free PMC article.
-
Nuclear IL-33 restrains the early conversion of fibroblasts to an extracellular matrix-secreting phenotype.Sci Rep. 2021 Jan 8;11(1):108. doi: 10.1038/s41598-020-80509-5. Sci Rep. 2021. PMID: 33420328 Free PMC article.
-
Quantitative model for inferring dynamic regulation of the tumour suppressor gene p53.BMC Bioinformatics. 2010 Jan 19;11:36. doi: 10.1186/1471-2105-11-36. BMC Bioinformatics. 2010. PMID: 20085646 Free PMC article.
-
Microarray data analysis and mining tools.Bioinformation. 2011 Apr 22;6(3):95-9. doi: 10.6026/97320630006095. Bioinformation. 2011. PMID: 21584183 Free PMC article.
-
Classification algorithms for phenotype prediction in genomics and proteomics.Front Biosci. 2008 Jan 1;13:691-708. doi: 10.2741/2712. Front Biosci. 2008. PMID: 17981580 Free PMC article. Review.
References
-
- Alizadeh AA, Eisen MB, Davis RE, Ma C, Lossos IS, Rosenwald A, Boldrick JC, Sabet H, Tran T, Yu X, Powell JI, Yang L, Marti GE, Moore T, Hudson J, Jr, Lu L, Lewis DB, Tibshirani R, Sherlock G, Chan WC, Greiner TC, Weisenburger DD, Armitage JO, Warnke R, Staudt LM. Distinct types of diffuse large B-cell lymphoma identified by gene expression profiling. Nature. 2002;403:503–511. - PubMed
-
- Bezdek JC, Pal SK. Fuzzy models for pattern recognition method that search for structures in data. IEEE press New York. 1992.
-
- Ben-Dor A, Bruhn L, Friedman N, Nachman I, Washington U. Tissue classification with gene expression profiles. RECOMB Tokyo Japan. 2000. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

