The SARS-CoV S glycoprotein: expression and functional characterization
- PMID: 14651994
- PMCID: PMC7111010
- DOI: 10.1016/j.bbrc.2003.11.054
The SARS-CoV S glycoprotein: expression and functional characterization
Abstract
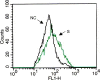
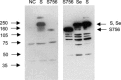
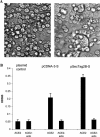
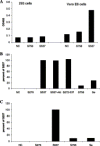
We have cloned, expressed, and characterized the full-length and various soluble fragments of the SARS-CoV (Tor2 isolate) S glycoprotein. Cells expressing S fused with receptor-expressing cells at neutral pH suggesting that the recombinant glycoprotein is functional, its membrane fusogenic activity does not require other viral proteins, and that low pH is not required for triggering membrane fusion; fusion was not observed at low receptor concentrations. S and its soluble ectodomain, S(e), were not cleaved to any significant degree. They ran at about 180-200kDa in SDS gels suggesting post-translational modifications as predicted by previous computer analysis and observed for other coronaviruses. Fragments containing the N-terminal amino acid residues 17-537 and 272-537 but not 17-276 bound specifically to Vero E6 cells and purified soluble receptor, ACE2, recently identified by M. Farzan and co-workers [Nature 426 (2003) 450-454]. Together with data for inhibition of binding by antibodies developed against peptides from S, these findings suggest that the receptor-binding domain is located between amino acid residues 303 and 537. These results also confirm that ACE2 is a functional receptor for the SARS virus and may help in the elucidation of the mechanisms of SARS-CoV entry and in the development of vaccine immunogens and entry inhibitors.
Figures


References
-
- Dimitrov D.S. Cell biology of virus entry. Cell. 2000;101:697–702. - PubMed
-
- Holmes K.V. SARS-associated coronavirus. N. Engl. J. Med. 2003;348:1948–1951. - PubMed
-
- Ksiazek T.G., Erdman D., Goldsmith C.S., Zaki S.R., Peret T., Emery S., Tong S., Urbani C., Comer J.A., Lim W., Rollin P.E., Dowell S.F., Ling A.E., Humphrey C.D., Shieh W.J., Guarner J., Paddock C.D., Rota P., Fields B., DeRisi J., Yang J.Y., Cox N., Hughes J.M., LeDuc J.W., Bellini W.J., Anderson L.J. A novel coronavirus associated with severe acute respiratory syndrome. N. Engl. J. Med. 2003;348:1953–1966. - PubMed
-
- Rota P.A., Oberste M.S., Monroe S.S., Nix W.A., Campagnoli R., Icenogle J.P., Penaranda S., Bankamp B., Maher K., Chen M.H., Tong S., Tamin A., Lowe L., Frace M., DeRisi J.L., Chen Q., Wang D., Erdman D.D., Peret T.C., Burns C., Ksiazek T.G., Rollin P.E., Sanchez A., Liffick S., Holloway B., Limor J., McCaustland K., Olsen-Rasmussen M., Fouchier R., Gunther S., Osterhaus A.D., Drosten C., Pallansch M.A., Anderson L.J., Bellini W.J. Characterization of a novel coronavirus associated with severe acute respiratory syndrome. Science. 2003;300:1394–1399. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

