Protein identification from two-dimensional gel electrophoresis analysis of Klebsiella pneumoniae by combined use of mass spectrometry data and raw genome sequences
- PMID: 14653859
- PMCID: PMC317362
- DOI: 10.1186/1477-5956-1-6
Protein identification from two-dimensional gel electrophoresis analysis of Klebsiella pneumoniae by combined use of mass spectrometry data and raw genome sequences
Abstract
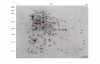
Separation of proteins by two-dimensional gel electrophoresis (2-DE) coupled with identification of proteins through peptide mass fingerprinting (PMF) by matrix-assisted laser desorption ionization time-of-flight mass spectrometry (MALDI-TOF MS) is the widely used technique for proteomic analysis. This approach relies, however, on the presence of the proteins studied in public-accessible protein databases or the availability of annotated genome sequences of an organism. In this work, we investigated the reliability of using raw genome sequences for identifying proteins by PMF without the need of additional information such as amino acid sequences. The method is demonstrated for proteomic analysis of Klebsiella pneumoniae grown anaerobically on glycerol. For 197 spots excised from 2-DE gels and submitted for mass spectrometric analysis 164 spots were clearly identified as 122 individual proteins. 95% of the 164 spots can be successfully identified merely by using peptide mass fingerprints and a strain-specific protein database (ProtKpn) constructed from the raw genome sequences of K. pneumoniae. Cross-species protein searching in the public databases mainly resulted in the identification of 57% of the 66 high expressed protein spots in comparison to 97% by using the ProtKpn database. 10 dha regulon related proteins that are essential for the initial enzymatic steps of anaerobic glycerol metabolism were successfully identified using the ProtKpn database, whereas none of them could be identified by cross-species searching. In conclusion, the use of strain-specific protein database constructed from raw genome sequences makes it possible to reliably identify most of the proteins from 2-DE analysis simply through peptide mass fingerprinting.
Figures
Similar articles
-
Proteome analysis of hairy root from Panax ginseng C.A. Meyer using peptide fingerprinting, internal sequencing and expressed sequence tag data.Proteomics. 2003 Dec;3(12):2379-92. doi: 10.1002/pmic.200300619. Proteomics. 2003. PMID: 14673788
-
Strategic proteome analysis of Candida magnoliae with an unsequenced genome.Proteomics. 2004 Nov;4(11):3588-99. doi: 10.1002/pmic.200400966. Proteomics. 2004. PMID: 15378735
-
Development and Preliminary Application of a Peptide Mass Fingerprinting Technique in Proteome Research.Sheng Wu Hua Xue Yu Sheng Wu Wu Li Xue Bao (Shanghai). 2000;32(4):373-378. Sheng Wu Hua Xue Yu Sheng Wu Wu Li Xue Bao (Shanghai). 2000. PMID: 12075426
-
Proteomic profiling and protein identification by MALDI-TOF mass spectrometry in unsequenced parasitic nematodes.PLoS One. 2012;7(3):e33590. doi: 10.1371/journal.pone.0033590. Epub 2012 Mar 29. PLoS One. 2012. PMID: 22479418 Free PMC article.
-
Analysing proteomic data.Int J Parasitol. 2005 Apr 30;35(5):543-53. doi: 10.1016/j.ijpara.2005.01.013. Int J Parasitol. 2005. PMID: 15826646 Review.
Cited by
-
Global proteome response of Escherichia coli BL21 to production of human basic fibroblast growth factor in complex and defined medium.Eng Life Sci. 2017 Jun 20;17(8):881-891. doi: 10.1002/elsc.201700036. eCollection 2017 Aug. Eng Life Sci. 2017. PMID: 32624836 Free PMC article.
-
Identification of Outer Membrane and Exoproteins of Carbapenem-Resistant Multilocus Sequence Type 258 Klebsiella pneumoniae.PLoS One. 2015 Apr 20;10(4):e0123219. doi: 10.1371/journal.pone.0123219. eCollection 2015. PLoS One. 2015. PMID: 25893665 Free PMC article.
-
The intra- and extracellular proteome of Aspergillus niger growing on defined medium with xylose or maltose as carbon substrate.Microb Cell Fact. 2010 Apr 20;9:23. doi: 10.1186/1475-2859-9-23. Microb Cell Fact. 2010. PMID: 20406453 Free PMC article.
-
The metabolic potential of Escherichia coli BL21 in defined and rich medium.Microb Cell Fact. 2014 Mar 23;13(1):45. doi: 10.1186/1475-2859-13-45. Microb Cell Fact. 2014. PMID: 24656150 Free PMC article.
-
Comparative proteomic analysis of high cell density cultivations with two recombinant Bacillus megaterium strains for the production of a heterologous dextransucrase.Proteome Sci. 2006 Oct 5;4:19. doi: 10.1186/1477-5956-4-19. Proteome Sci. 2006. PMID: 17022804 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

