A novel allele of fission yeast rad11 that causes defects in DNA repair and telomere length regulation
- PMID: 14654689
- PMCID: PMC291861
- DOI: 10.1093/nar/gkg917
A novel allele of fission yeast rad11 that causes defects in DNA repair and telomere length regulation
Abstract
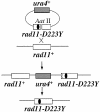
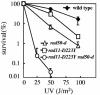
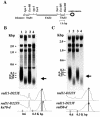
Replication protein A (RPA) is a heterotrimeric single-stranded DNA-binding protein involved in DNA replication, recombination and repair. In Saccharomyces cerevisiae, several mutants in the RFA1 gene encoding the large subunit of RPA have been isolated and one of the mutants with a missense allele, rfa1-D228Y, shows a synergistic reduction in telomere length when combined with a yku70 mutation. So far, only one mutant allele of the rad11(+) gene encoding the large subunit of RPA has been reported in Schizosaccharomyces pombe. To study the role of S.pombe RPA in DNA repair and possibly in telomere maintenance, we constructed a rad11-D223Y mutant, which corresponds to the S.cerevisiae rfa1-D228Y mutant. rad11-D223Y cells were methylmethane sulfonate, hydroxyurea, UV and gamma-ray sensitive, suggesting that rad11-D223Y cells have a defect in DNA repair activity. Unlike the S.cerevisiae rfa1-D228Y mutation, the rad11-D223Y mutation itself caused telomere shortening. Moreover, Rad11-Myc bound to telomere in a ChIP assay. These results strongly suggest that RPA is directly involved in telomere maintenance.
Figures
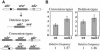



Similar articles
-
Characterization of genetic interactions with RFA1: the role of RPA in DNA replication and telomere maintenance.Biochimie. 2000 Jan;82(1):71-8. doi: 10.1016/s0300-9084(00)00183-8. Biochimie. 2000. PMID: 10717390
-
Molecular characterization of the Schizosaccharomyces pombe nbs1+ gene involved in DNA repair and telomere maintenance.Mol Cell Biol. 2003 Sep;23(18):6553-63. doi: 10.1128/MCB.23.18.6553-6563.2003. Mol Cell Biol. 2003. PMID: 12944481 Free PMC article.
-
The Schizosaccharomyces pombe rad11+ gene encodes the large subunit of replication protein A.Mol Cell Biol. 1997 May;17(5):2381-90. doi: 10.1128/MCB.17.5.2381. Mol Cell Biol. 1997. PMID: 9111307 Free PMC article.
-
Repair of UV damage in the fission yeast Schizosaccharomyces pombe.Mutat Res. 2000 Jun 30;451(1-2):197-210. doi: 10.1016/s0027-5107(00)00050-6. Mutat Res. 2000. PMID: 10915873 Review.
-
Molecular biology of DNA repair in the fission yeast Schizosaccharomyces pombe.Mutat Res. 1996 Aug 8;363(3):147-61. doi: 10.1016/0921-8777(96)00017-1. Mutat Res. 1996. PMID: 8765156 Review. No abstract available.
Cited by
-
RPA and POT1: friends or foes at telomeres?Cell Cycle. 2012 Feb 15;11(4):652-7. doi: 10.4161/cc.11.4.19061. Cell Cycle. 2012. PMID: 22373525 Free PMC article.
-
Gain-of-function mutations in RPA1 cause a syndrome with short telomeres and somatic genetic rescue.Blood. 2022 Feb 17;139(7):1039-1051. doi: 10.1182/blood.2021011980. Blood. 2022. PMID: 34767620 Free PMC article.
-
Chl1 coordinates with H3K9 methyltransferase Clr4 to reduce the accumulation of RNA-DNA hybrids and maintain genome stability.iScience. 2022 Apr 27;25(5):104313. doi: 10.1016/j.isci.2022.104313. eCollection 2022 May 20. iScience. 2022. PMID: 35602970 Free PMC article.
-
The Pif1 family helicase Pfh1 facilitates telomere replication and has an RPA-dependent role during telomere lengthening.DNA Repair (Amst). 2014 Dec;24:80-86. doi: 10.1016/j.dnarep.2014.09.008. Epub 2014 Oct 7. DNA Repair (Amst). 2014. PMID: 25303777 Free PMC article.
-
Smc5/6 Complex Promotes Rad3ATR Checkpoint Signaling at the Perturbed Replication Fork through Sumoylation of the RecQ Helicase Rqh1.Mol Cell Biol. 2022 Jun 16;42(6):e0004522. doi: 10.1128/mcb.00045-22. Epub 2022 May 25. Mol Cell Biol. 2022. PMID: 35612306 Free PMC article.
References
-
- Wold M.S. (1997) Replication protein A: a heterotrimeric, single-stranded DNA-binding protein required for eukaryotic DNA metabolism. Annu. Rev. Biochem., 66, 61–92. - PubMed
-
- Stillman B. (1989) Initiation of eukaryotic DNA replication in vitro. Annu. Rev. Cell Biol., 5, 197–245. - PubMed
-
- Coverley D., Kenny,M.K., Munn,M., Rupp,W.D., Lane,D.P. and Wood,R.D. (1991) Requirement for the replication protein SSB in human DNA excision repair. Nature, 349, 538–541. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

