Efficient degradation of 2,4,6-Trichlorophenol requires a set of catabolic genes related to tcp genes from Ralstonia eutropha JMP134(pJP4)
- PMID: 14660355
- PMCID: PMC309972
- DOI: 10.1128/AEM.69.12.7108-7115.2003
Efficient degradation of 2,4,6-Trichlorophenol requires a set of catabolic genes related to tcp genes from Ralstonia eutropha JMP134(pJP4)
Abstract
2,4,6-Trichlorophenol (2,4,6-TCP) is a hazardous pollutant. Several aerobic bacteria are known to degrade this compound. One of these, Ralstonia eutropha JMP134(pJP4), a well-known, versatile chloroaromatic compound degrader, is able to grow in 2,4,6-TCP by converting it to 2,6-dichlorohydroquinone, 6-chlorohydroxyquinol, 2-chloromaleylacetate, maleylacetate, and beta-ketoadipate. Three enzyme activities encoded by tcp genes, 2,4,6-TCP monooxygenase (tcpA), 6-chlorohydroxyquinol 1,2-dioxygenase (tcpC), and maleylacetate reductase (tcpD), are involved in this catabolic pathway. Here we provide evidence that all these tcp genes are clustered in the R. eutropha JMP134(pJP4) chromosome, forming the putative catabolic operon tcpRXABCYD. We studied the presence of tcp-like gene sequences in several other 2,4,6-TCP-degrading bacterial strains and found two types of strains. One type includes strains belonging to the Ralstonia genus and possessing a set of tcp-like genes, which efficiently degrade 2,4,6-TCP and therefore grow in liquid cultures containing this chlorophenol as a sole carbon source. The other type includes strains belonging to the genera Pseudomonas, Sphingomonas, or Sphingopixis, which do not have tcp-like gene sequences and degrade this pollutant less efficiently and which therefore grow only as small colonies on plates with 2,4,6-TCP. Other than strain JMP134, none of the bacterial strains whose genomes have been sequenced possesses a full set of tcp-like gene sequences.
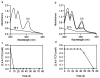
Figures


Similar articles
-
Genetic characterization of 2,4,6-trichlorophenol degradation in Cupriavidus necator JMP134.Appl Environ Microbiol. 2007 May;73(9):2769-76. doi: 10.1128/AEM.02584-06. Epub 2007 Feb 23. Appl Environ Microbiol. 2007. PMID: 17322325 Free PMC article.
-
Genetic and biochemical characterization of a 2,4,6-trichlorophenol degradation pathway in Ralstonia eutropha JMP134.J Bacteriol. 2002 Jul;184(13):3492-500. doi: 10.1128/JB.184.13.3492-3500.2002. J Bacteriol. 2002. PMID: 12057943 Free PMC article.
-
Degradation of 2,4,6-trichlorophenol via chlorohydroxyquinol in Ralstonia eutropha JMP134 and JMP222.J Basic Microbiol. 2000;40(4):243-9. doi: 10.1002/1521-4028(200008)40:4<243::AID-JOBM243>3.0.CO;2-D. J Basic Microbiol. 2000. PMID: 10986670
-
Metabolic reconstruction of aromatic compounds degradation from the genome of the amazing pollutant-degrading bacterium Cupriavidus necator JMP134.FEMS Microbiol Rev. 2008 Aug;32(5):736-94. doi: 10.1111/j.1574-6976.2008.00122.x. Epub 2008 Aug 7. FEMS Microbiol Rev. 2008. PMID: 18691224 Review.
-
Microbial degradation of 2,4-dichlorophenoxyacetic acid: Insight into the enzymes and catabolic genes involved, their regulation and biotechnological implications.Crit Rev Microbiol. 2016;42(2):194-208. doi: 10.3109/1040841X.2014.917068. Epub 2014 Jul 24. Crit Rev Microbiol. 2016. PMID: 25058513 Review.
Cited by
-
A previously unexposed forest soil microbial community degrades high levels of the pollutant 2,4,6-trichlorophenol.Appl Environ Microbiol. 2004 Dec;70(12):7567-70. doi: 10.1128/AEM.70.12.7567-7570.2004. Appl Environ Microbiol. 2004. PMID: 15574963 Free PMC article.
-
Novel gene clusters and metabolic pathway involved in 3,5,6-trichloro-2-pyridinol degradation by Ralstonia sp. strain T6.Appl Environ Microbiol. 2013 Dec;79(23):7445-53. doi: 10.1128/AEM.01817-13. Epub 2013 Sep 20. Appl Environ Microbiol. 2013. PMID: 24056464 Free PMC article.
-
Genetic characterization of 2,4,6-trichlorophenol degradation in Cupriavidus necator JMP134.Appl Environ Microbiol. 2007 May;73(9):2769-76. doi: 10.1128/AEM.02584-06. Epub 2007 Feb 23. Appl Environ Microbiol. 2007. PMID: 17322325 Free PMC article.
-
Oxidative dehalogenation and denitration by a flavin-dependent monooxygenase is controlled by substrate deprotonation.Chem Sci. 2018 Aug 8;9(38):7468-7482. doi: 10.1039/c8sc01482e. eCollection 2018 Oct 14. Chem Sci. 2018. PMID: 30319747 Free PMC article.
-
Survival of prokaryotes in a polluted waste dump during remediation by alkaline hydrolysis.Ecotoxicology. 2014 Apr;23(3):404-18. doi: 10.1007/s10646-014-1205-y. Epub 2014 Feb 16. Ecotoxicology. 2014. PMID: 24532314
References
-
- Andreoni, V., G. Baggi, M. Colombo, L. Cavalca, M. Zangrossi, and S. Bernasconi. 1998. Degradation of 2,4,6-trichlorophenol by a specialized organism and by indigenous soil microflora: bioaugmentation and self-remediability for soil restoration. Lett. Appl. Microbiol. 27:86-92. - PubMed
-
- Aranda, C., F. Godoy, B. González, J. Homo, and M. Martínez. 1999. Effects of glucose and phenylalanine upon 2,4,6-trichlorophenol degradation by Pseudomonas paucimobilis S37 cells in non-growth state. Microbios 100:73-82. - PubMed
-
- Ausubel, F., R. Brent, R. Kingston, D. Moore, J. Seidman, J. Smith, and K. Struhl. 1992. Short protocols in molecular biology: a compendium of methods from, current protocols in molecular biology, 2nd ed. John Wiley & Sons, Inc., New York, N.Y.
-
- Bock, C., R. M. Kroppenstedt, U. Schmidt, and H. Diekmann. 1996. Degradation of prochloraz and 2,4,6-trichlorophenol by environmental bacterial strains. Appl. Microbiol. Biotechnol. 45:257-262. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

