Genomes of the parapoxviruses ORF virus and bovine papular stomatitis virus
- PMID: 14671098
- PMCID: PMC303426
- DOI: 10.1128/jvi.78.1.168-177.2004
Genomes of the parapoxviruses ORF virus and bovine papular stomatitis virus
Abstract
Bovine papular stomatitis virus (BPSV) and orf virus (ORFV), members of the genus Parapoxvirus of the Poxviridae, are etiologic agents of worldwide diseases affecting cattle and small ruminants, respectively. Here we report the genomic sequences and comparative analysis of BPSV strain BV-AR02 and ORFV strains OV-SA00, isolated from a goat, and OV-IA82, isolated from a sheep. Parapoxvirus (PPV) BV-AR02, OV-SA00, and OV-IA82 genomes range in size from 134 to 139 kbp, with an average nucleotide composition of 64% G+C. BPSV and ORFV genomes contain 131 and 130 putative genes, respectively, and share colinearity over 127 genes, 88 of which are conserved in all characterized chordopoxviruses. BPSV and ORFV contain 15 and 16 open reading frames (ORFs), respectively, which lack similarity to other poxvirus or cellular proteins. All genes with putative roles in pathogenesis, including a vascular endothelial growth factor (VEGF)-like gene, are present in both viruses; however, BPSV contains two extra ankyrin repeat genes absent in ORFV. Interspecies sequence variability is observed in all functional classes of genes but is highest in putative virulence/host range genes, including genes unique to PPV. At the amino acid level, OV-SA00 is 94% identical to OV-IA82 and 71% identical to BV-AR02. Notably, ORFV 006/132, 103, 109, 110, and 116 genes (VEGF, homologues of vaccinia virus A26L, A33R, and A34R, and a novel PPV ORF) show an unusual degree of intraspecies variability. These genomic differences are consistent with the classification of BPSV and ORFV as two PPV species. Compared to other mammalian chordopoxviruses, PPV shares unique genomic features with molluscum contagiosum virus, including a G+C-rich nucleotide composition, three orthologous genes, and a paucity of nucleotide metabolism genes. Together, these data provide a comparative view of PPV genomics.
Figures
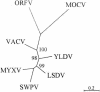
References
-
- Beattie, E., E. B. Kauffman, H. Martinez, M. E. Perkus, B. L. Jacobs, E. Paoletti, and J. Tartaglia. 1996. Host-range restriction of vaccinia virus E3L-specific deletion mutants. Virus Genes 12:89-94. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources

