The oxidized deoxynucleoside triphosphate pool is a significant contributor to genetic instability in mismatch repair-deficient cells
- PMID: 14673178
- PMCID: PMC303369
- DOI: 10.1128/MCB.24.1.465-474.2004
The oxidized deoxynucleoside triphosphate pool is a significant contributor to genetic instability in mismatch repair-deficient cells
Abstract
Oxidation is a common form of DNA damage to which purines are particularly susceptible. We previously reported that oxidized dGTP is potentially an important source of DNA 8-oxodGMP in mammalian cells and that the incorporated lesions are removed by DNA mismatch repair (MMR). MMR deficiency is associated with a mutator phenotype and widespread microsatellite instability (MSI). Here, we identify oxidized deoxynucleoside triphosphates (dNTPs) as an important cofactor in this genetic instability. The high spontaneous hprt mutation rate of MMR-defective msh2(-/-) mouse embryonic fibroblasts was attenuated by expression of the hMTH1 protein, which degrades oxidized purine dNTPs. A high level of hMTH1 abolished their mutator phenotype and restored the hprt mutation rate to normal. Molecular analysis of hprt mutants showed that the presence of hMTH1 reduced the incidence of mutations in all classes, including frameshifts, and also implicated incorporated 2-oxodAMP in the mutator phenotype. In hMSH6-deficient DLD-1 human colorectal carcinoma cells, overexpression of hMTH1 markedly attenuated the spontaneous mutation rate and reduced MSI. It also reduced the incidence of -G and -A frameshifts in the hMLH1-defective DU145 human prostatic cancer cell line. Our findings indicate that incorporation of oxidized purines from the dNTP pool may contribute significantly to the extreme genetic instability of MMR-defective human tumors.
Figures




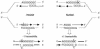
References
-
- Aaltonen, L. A., P. Peltomaki, F. S. Leach, P. Sistonen, L. Pylkkanen, J.-P. Mecklin, H. Jarvinen, S. M. Powell, J. Jen, S. R. Hamilton, G. M. Petersen, K. W. Kinzler, B. Vogelstein, and A. de la Chapelle. 1993. Clues to the pathogenesis of familial colorectal cancer. Science 260:812-816. - PubMed
-
- Akiyama, M., T. Horiuchi, and M. Sekiguchi. 1987. Molecular cloning and nucleotide sequence of the mutT mutator of Escherichia coli that causes A:T to C:G transversion. Mol. Gen. Genet. 206:9-16. - PubMed
-
- Andrew, S. E., A. H. Reitmair, J. Fox, L. Hsiao, A. Francis, M. McKinnon, T. W. Mak, and F. R. Jirik. 1997. Base transitions dominate the mutational spectrum of a transgenic reporter gene in MSH2 deficient mice. Oncogene 15:123-129. - PubMed
-
- Bhattacharyya, N., A. Ganesh, G. Phear, B. Richards, A. Skandalis, and M. Meuth. 1995. Molecular analysis of mutations in mutator colorectal carcinoma cell lines. Hum. Mol. Genet. 4:2057-2064. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous
