Ex vivo screening for immunodominant viral epitopes by quantitative real time polymerase chain reaction (qRT-PCR)
- PMID: 14675481
- PMCID: PMC324417
- DOI: 10.1186/1479-5876-1-12
Ex vivo screening for immunodominant viral epitopes by quantitative real time polymerase chain reaction (qRT-PCR)
Abstract
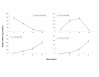
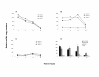
The identification and characterization of viral epitopes across the Human Leukocyte Antigen (HLA) polymorphism is critical for the development of actives-specific or adoptive immunotherapy of virally-mediated diseases. This work investigates whether cytokine mRNA transcripts could be used to identify epitope-specific HLA-restricted memory T lymphocytes reactivity directly in fresh peripheral blood mononuclear cells (PBMCs) from viral-seropositive individuals in response to ex vivo antigen recall. PBMCs from HLA-A*0201 healthy donors, seropositive for Cytomegalovirus (CMV) and Influenza (Flu), were exposed for different periods and at different cell concentrations to the HLA-A*0201-restricted viral FluM158-66 and CMVpp65495-503 peptides. Quantitative real time PCR (qRT-PCR) was employed to evaluate memory T lymphocyte immune reactivation by measuring the production of mRNA encoding four cytokines: Interferon-gamma (IFN-gamma), Interleukin-2 (IL-2), Interleukin-4 (IL-4), and Interleukin-10 (IL-10). We could characterize cytokine expression kinetics that illustrated how cytokine mRNA levels could be used as ex vivo indicators of T cell reactivity. Particularly, IFN-gamma mRNA transcripts could be consistently detected within 3 to 12 hours of short-term stimulation in levels sufficient to screen for HLA-restricted viral immune responses in seropositive subjects. This strategy will enhance the efficiency of the identification of viral epitopes independently of the individual HLA phenotype and could be used to follow the intensity of immune responses during disease progression or in response to in vivo antigen-specific immunization.
Figures



References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

