Multiple formate dehydrogenase enzymes in the facultative methylotroph Methylobacterium extorquens AM1 are dispensable for growth on methanol
- PMID: 14679220
- PMCID: PMC303455
- DOI: 10.1128/JB.186.1.22-28.2004
Multiple formate dehydrogenase enzymes in the facultative methylotroph Methylobacterium extorquens AM1 are dispensable for growth on methanol
Abstract
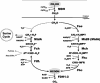
Formate dehydrogenase has traditionally been assumed to play an essential role in energy generation during growth on C(1) compounds. However, this assumption has not yet been experimentally tested in methylotrophic bacteria. In this study, a whole-genome analysis approach was used to identify three different formate dehydrogenase systems in the facultative methylotroph Methylobacterium extorquens AM1 whose expression is affected by either molybdenum or tungsten. A complete set of single, double, and triple mutants was generated, and their phenotypes were analyzed. The growth phenotypes of the mutants suggest that any one of the three formate dehydrogenases is sufficient to sustain growth of M. extorquens AM1 on formate, while surprisingly, none is required for growth on methanol or methylamine. Nuclear magnetic resonance analysis of the fate of [(13)C]methanol revealed that while cells of wild-type M. extorquens AM1 as well as cells of all the single and the double mutants continuously produced [(13)C]bicarbonate and (13)CO(2), cells of the triple mutant accumulated [(13)C]formate instead. Further studies of the triple mutant showed that formate was not produced quantitatively and was consumed later in growth. These results demonstrated that all three formate dehydrogenase systems must be inactivated in order to disrupt the formate-oxidizing capacity of the organism but that an alternative formate-consuming capacity exists in the triple mutant.
Figures


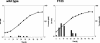
References
-
- Anthony, C. 1982. The biochemistry of methylotrophs. Academic Press, New York, N.Y.
-
- Anthony, C., and P. Williams. 2003. The structure and mechanism of methanol dehydrogenase. Biochim. Biophys. Acta 1647:18-23. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

