Characterization of the RNA components of a putative molecular switch in the 3' untranslated region of the murine coronavirus genome
- PMID: 14694098
- PMCID: PMC368785
- DOI: 10.1128/jvi.78.2.669-682.2004
Characterization of the RNA components of a putative molecular switch in the 3' untranslated region of the murine coronavirus genome
Abstract
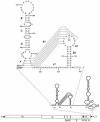
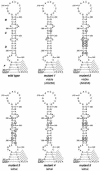
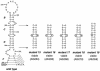
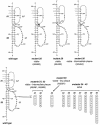
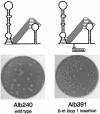
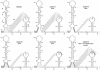
RNA virus genomes contain cis-acting sequence and structural elements that participate in viral replication. We previously identified a bulged stem-loop secondary structure at the upstream end of the 3' untranslated region (3' UTR) of the genome of the coronavirus mouse hepatitis virus (MHV). This element, beginning immediately downstream of the nucleocapsid gene stop codon, was shown to be essential for virus replication. Other investigators discovered an adjacent downstream pseudoknot in the 3' UTR of the closely related bovine coronavirus (BCoV). This pseudoknot was also shown to be essential for replication, and it has a conserved counterpart in every group 1 and group 2 coronavirus. In MHV and BCoV, the bulged stem-loop and pseudoknot are, in part, mutually exclusive, because of the overlap of the last segment of the stem-loop and stem 1 of the pseudoknot. This led us to hypothesize that they form a molecular switch, possibly regulating a transition occurring during viral RNA synthesis. We have now performed an extensive genetic analysis of the two components of this proposed switch. Our results define essential and nonessential components of these structures and establish the limits to which essential parts of each element can be destabilized prior to loss of function. Most notably, we have confirmed the interrelationship of the two putative switch elements. Additionally, we have identified a pseudoknot loop insertion mutation that appears to point to a genetic interaction between the pseudoknot and a distant region of the genome.
Figures




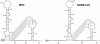
References
-
- Horton, R. M., and L. R. Pease. 1991. Recombination and mutagenesis of DNA sequences using PCR, p. 217-247. In M. J. McPherson (ed.), Directed mutagenesis, a practical approach. IRL Press, New York, N.Y.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

