Whole genome amplification of DNA from laser capture-microdissected tissue for high-throughput single nucleotide polymorphism and short tandem repeat genotyping
- PMID: 14695315
- PMCID: PMC1602222
- DOI: 10.1016/S0002-9440(10)63092-1
Whole genome amplification of DNA from laser capture-microdissected tissue for high-throughput single nucleotide polymorphism and short tandem repeat genotyping
Abstract
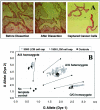
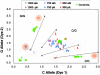
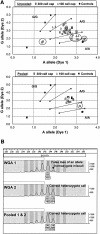
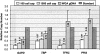
Genome-wide screening of genetic alterations between normal and cancer cells, as well as among subgroups of tumors, is important for establishing molecular mechanism and classification of cancer. Gene silencing through loss of heterozygosity is widely observed in cancer cells and detectable by analyzing allelic loss of single nucleotide polymorphism and/or short tandem repeat markers. To use minute quantities of DNA that are available through laser capture microdissection (LCM) of cancer cells, a whole genome amplification method that maintains locus and allele balance is essential. We have successfully used a ø29 polymerase-based isothermal whole genome amplification method to amplify LCM DNA using a proteinase K lysis procedure coupled with a pooling strategy. Through single nucleotide polymorphism and short tandem repeat genotype analysis we demonstrate that using pooled DNA from two or three separate amplification reactions significantly reduces any allele bias introduced during amplification. This strategy is especially effective when using small quantities of source DNA. Although a convenient alkaline lysis DNA extraction procedure provided satisfactory results from using 1500 to 3000 LCM cells, proteinase K digestion was superior for lower cell numbers. Accurate genotyping is achieved with as few as 100 cells when both proteinase K extraction and pooling are applied.
Figures

References
-
- Weber BL. Cancer genomics. Cancer Cell. 2002;1:37–47. - PubMed
-
- Rosenwald A, Wright G, Chan WC, Connors JM, Campo E, Fisher RI, Gascoyne RD, Muller-Hermelink HK, Smeland EB, Giltnane JM, Hurt EM, Zhao H, Averett L, Yang L, Wilson WH, Jaffe ES, Simon R, Klausner RD, Powell J, Duffey PL, Longo DL, Greiner TC, Weisenburger DD, Sanger WG, Dave BJ, Lynch JC, Vose J, Armitage JO, Montserrat E, Lopez-Guillermo A, Grogan TM, Miller TP, LeBlanc M, Ott G, Kvaloy S, Delabie J, Holte H, Krajci P, Stokke T, Staudt LM. Lymphoma/Leukemia Molecular Profiling Project: The use of molecular profiling to predict survival after chemotherapy for diffuse large-B-cell lymphoma. N Engl J Med. 2002;346:1937–1947. - PubMed
-
- van ’t Veer LJ, Dai H, van de Vijver MJ, He YD, Hart AA, Mao M, Peterse HL, van der Kooy K, Marton MJ, Witteveen AT, Schreiber GJ, Kerkhoven RM, Roberts C, Linsley PS, Bernards R, Friend SH. Gene expression profiling predicts clinical outcome of breast cancer. Nature. 2002;415:530–536. - PubMed
-
- Ramaswamy S, Ross KN, Lander ES, Golub TR. A molecular signature of metastasis in primary solid tumors. Nat Genet. 2003;33:49–54. - PubMed
-
- Thiagalingam S, Foy RL, Cheng KH, Lee HJ, Thiagalingam A, Ponte JF. Loss of heterozygosity as a predictor to map tumor suppressor genes in cancer: molecular basis of its occurrence. Curr Opin Oncol. 2002;14:65–72. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

