Meiotic pairing and imprinted X chromatin assembly in Caenorhabditis elegans
- PMID: 14702046
- PMCID: PMC4098868
- DOI: 10.1038/ng1283
Meiotic pairing and imprinted X chromatin assembly in Caenorhabditis elegans
Abstract
The genetic imprinting of individual loci or whole chromosomes, as in imprinted X-chromosome inactivation in mammals, is established and reset during gametogenesis; defects in this process in the parent can result in disease in the offspring. We describe a sperm-specific chromatin-based imprinting of the X chromosome in the nematode Caenorhabditis elegans that is restricted to histone H3 modifications. The epigenetic imprint is established during spermatogenesis and its stability in the offspring is affected by the presence of a pairing partner during meiosis in the parental germ line. We observed that DNA lacking a pairing partner during meiosis, the normal situation for the X chromosome in males, is targeted for methylation of histone H3 at Lys9 (H3-Lys9) and can be silenced. Targeting unpaired DNA for silencing during meiosis, a potential hallmark of genome defense, could therefore have a conserved role in imprinted X-chromosome inactivation and, ultimately, in sex chromosome evolution.
Conflict of interest statement
The authors declare that they have no competing financial interests.
Figures
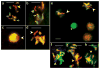
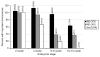

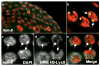
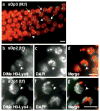
Comment in
-
Marking Xs, together and separately.Nat Genet. 2004 Jan;36(1):12-3. doi: 10.1038/ng0104-12. Nat Genet. 2004. PMID: 14702033 No abstract available.
References
-
- Huynh KD, Lee JT. Imprinted X inactivation in eutherians: a model of gametic execution and zygotic relaxation. Curr Opin Cell Biol. 2001;13:690–697. - PubMed
-
- Reik W, Walter J. Genomic imprinting: parental influence on the genome. Nat Rev Genet. 2001;2:21–32. - PubMed
-
- Heard E, et al. Methylation of histone H3 at Lys-9 is an early mark on the X chromosome during X inactivation. Cell. 2001;107:727–738. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

