Occurrence and characterization of mercury resistance in the hyperthermophilic archaeon Sulfolobus solfataricus by use of gene disruption
- PMID: 14702312
- PMCID: PMC305765
- DOI: 10.1128/JB.186.2.427-437.2004
Occurrence and characterization of mercury resistance in the hyperthermophilic archaeon Sulfolobus solfataricus by use of gene disruption
Abstract
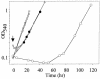
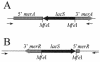
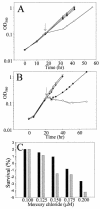
Mercury resistance mediated by mercuric reductase (MerA) is widespread among bacteria and operates under the control of MerR. MerR represents a unique class of transcription factors that exert both positive and negative regulation on gene expression. Archaea and bacteria are prokaryotes, yet little is known about the biological role of mercury in archaea or whether a resistance mechanism occurs in these organisms. The archaeon Sulfolobus solfataricus was sensitive to mercuric chloride, and low-level adaptive resistance could be induced by metal preconditioning. Protein phylogenetic analysis of open reading frames SSO2689 and SSO2688 clarified their identity as orthologs of MerA and MerR. Northern analysis established that merA transcription responded to mercury challenge, since mRNA levels were transiently induced and, when normalized to 7S RNA, approximated values for other highly expressed transcripts. Primer extension analysis of merA mRNA predicted a noncanonical TATA box with nonstandard transcription start site spacing. The functional roles of merA and merR were clarified further by gene disruption. The merA mutant exhibited mercury sensitivity relative to wild type and was defective in elemental mercury volatilization, while the merR mutant was mercury resistant. Northern analysis of the merR mutant revealed merA transcription was constitutive and that transcript abundance was at maximum levels. These findings constitute the first report of an archaeal heavy metal resistance system; however, unlike bacteria the level of resistance is much lower. The archaeal system employs a divergent MerR protein that acts only as a negative transcriptional regulator of merA expression.
Figures
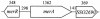




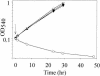

References
-
- Allen, M. B. 1959. Studies with Cyanidium caldarium, an anomalously pigmented chlorophyte. Arch. Mikrobiol. 32:270-277. - PubMed
-
- Ansari, A. Z., M. L. Chael, and T. V. O'Halloran. 1992. Allosteric underwinding of DNA is a critical step in positive control of transcription by Hg-MerR. Nature 355:87-89. - PubMed
-
- Ansari, A. Z., J. E. Bradner, and T. V. O'Halloran. 1995. DNA-bend modulation in a repressor-to-activator switching mechanism. Nature 374:371-375. - PubMed
-
- Barkay, T., S. M. Miller, and A. O. Summers. 2003. Bacterial mercury resistance from atoms to ecosystems. FEMS Microbiol. Rev. 27:355-384. - PubMed
-
- Baumann, P., S. A. Qureshi, and S. P. Jackson. 1995. Transcription: new insights from studies on Archaea. Trends Genet. 11:279-283. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

