RNAi-mediated targeting of heterochromatin by the RITS complex
- PMID: 14704433
- PMCID: PMC3244756
- DOI: 10.1126/science.1093686
RNAi-mediated targeting of heterochromatin by the RITS complex
Erratum in
- Science. 2013 Oct 11;342(6155):193
Abstract
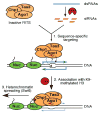
RNA interference (RNAi) is a widespread silencing mechanism that acts at both the posttranscriptional and transcriptional levels. Here, we describe the purification of an RNAi effector complex termed RITS (RNA-induced initiation of transcriptional gene silencing) that is required for heterochromatin assembly in fission yeast. The RITS complex contains Ago1 (the fission yeast Argonaute homolog), Chp1 (a heterochromatin-associated chromodomain protein), and Tas3 (a novel protein). In addition, the complex contains small RNAs that require the Dicer ribonuclease for their production. These small RNAs are homologous to centromeric repeats and are required for the localization of RITS to heterochromatic domains. The results suggest a mechanism for the role of the RNAi machinery and small RNAs in targeting of heterochromatin complexes and epigenetic gene silencing at specific chromosomal loci.
Figures




References
-
- Grewal SIS. J Cell Physiol. 2000;184:311. - PubMed
-
- Grewal SIS, Moazed D. Science. 2003;301:798. - PubMed
-
- Partridge JF, Scott KS, Bannister AJ, Kouzarides T, Allshire RC. Curr Biol. 2002;12:1652. - PubMed
-
- Zilberman D, Cao X, Jacobsen SE. Science. 2003;299:716. - PubMed
-
- Matzke M, Matzke AJM, Kooter JM. Science. 2001;293:1080. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

