Schnurri-3 (KRC) interacts with c-Jun to regulate the IL-2 gene in T cells
- PMID: 14707112
- PMCID: PMC1887724
- DOI: 10.1084/jem.20030421
Schnurri-3 (KRC) interacts with c-Jun to regulate the IL-2 gene in T cells
Abstract
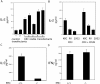
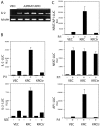
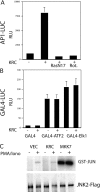
The activator protein 1 (AP-1) transcription factor is a key participant in the control of T cell proliferation, cytokine production, and effector function. In the immune system, AP-1 activity is highest in T cells, suggesting that a subset of T cell-specific coactivator proteins exist to selectively potentiate AP-1 function. Here, we describe that the expression of Schnurri-3, also known as kappa recognition component (KRC), is induced upon T cell receptor signaling in T cells and functions to regulate the expression of the interleukin 2 (IL-2) gene. Overexpression of KRC in transformed and primary T cells leads to increased IL-2 production, whereas dominant-negative KRC, or loss of KRC protein in KRC-null mice, results in diminished IL-2 production. KRC physically associates with the c-Jun transcription factor and serves as a coactivator to augment AP-1-dependent IL-2 gene transcription.
Figures

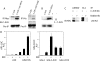
Similar articles
-
Regulation of nuclear factor-kappa B and activator protein-1 activities after stimulation of T cells via glycosylphosphatidylinositol-anchored Ly-6A/E.J Immunol. 1994 Sep 15;153(6):2394-406. J Immunol. 1994. PMID: 7915738
-
Involvement of Rel, Fos, and Jun proteins in binding activity to the IL-2 promoter CD28 response element/AP-1 sequence in human T cells.J Immunol. 1997 Aug 1;159(3):1319-27. J Immunol. 1997. PMID: 9233628
-
Altered AP-1 (activating protein-1) activity and c-jun activation in T cells exposed to the amide class herbicide 3,4-dichloropropionanilide (DCPA).Toxicol Sci. 2004 May;79(1):98-105. doi: 10.1093/toxsci/kfh090. Epub 2004 Feb 19. Toxicol Sci. 2004. PMID: 14976341
-
Regulation of bone formation and immune cell development by Schnurri proteins.Adv Exp Med Biol. 2010;658:117-22. doi: 10.1007/978-1-4419-1050-9_13. Adv Exp Med Biol. 2010. PMID: 19950022 Review.
-
Fluorescence-based monitoring of interleukin-2 gene expression.Immunol Rev. 1991 Feb;119:95-103. doi: 10.1111/j.1600-065x.1991.tb00579.x. Immunol Rev. 1991. PMID: 1904398 Review. No abstract available.
Cited by
-
Disruption of ZAS3 in mice alters NF-kappaB and AP-1 DNA binding and T-cell development.Gene Expr. 2007;14(2):83-100. doi: 10.3727/105221607783417574. Gene Expr. 2007. PMID: 18257392 Free PMC article.
-
Identification and characterization of extrachromosomal circular DNA in large-artery atherosclerotic stroke.J Cell Mol Med. 2024 Apr;28(7):e18210. doi: 10.1111/jcmm.18210. J Cell Mol Med. 2024. PMID: 38506071 Free PMC article.
-
Posttranscriptional deregulation of Src due to aberrant miR34a and miR203 contributes to gastric cancer development.BMB Rep. 2013 Jun;46(6):316-21. doi: 10.5483/bmbrep.2013.46.6.208. BMB Rep. 2013. PMID: 23790975 Free PMC article.
-
The large zinc finger protein ZAS3 is a critical modulator of osteoclastogenesis.PLoS One. 2011 Mar 3;6(3):e17161. doi: 10.1371/journal.pone.0017161. PLoS One. 2011. PMID: 21390242 Free PMC article.
-
Schnurri-3 regulates BMP9-induced osteogenic differentiation and angiogenesis of human amniotic mesenchymal stem cells through Runx2 and VEGF.Cell Death Dis. 2020 Jan 29;11(1):72. doi: 10.1038/s41419-020-2279-5. Cell Death Dis. 2020. Retraction in: Cell Death Dis. 2021 Dec 9;12(12):1144. doi: 10.1038/s41419-021-04443-8. PMID: 31996667 Free PMC article. Retracted.
References
-
- Wu, L.C., Y. Liu, J. Strandtmann, C.H. Mak, B. Lee, Z. Li, and C.Y. Yu. 1996. The mouse DNA binding protein Rc for the kappa B motif of transcription and for the V(D)J recombination signal sequences contains composite DNA-protein interaction domains and belongs to a new family of large transcriptional proteins. Genomics. 35:415–424. - PubMed
-
- Hicar, M.D., M.L. Robinson, and L.C. Wu. 2002. Embryonic expression and regulation of the large zinc finger protein KRC. Genesis. 33:8–20. - PubMed
-
- Mak, C.H., Z. Li, C.E. Allen, Y. Liu, and L. Wu. 1998. KRC transcripts: identification of an unusual alternative splicing event. Immunogenetics. 48:32–39. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

