Impact of DNA polymerases and their buffer systems on quantitative real-time PCR
- PMID: 14715792
- PMCID: PMC321720
- DOI: 10.1128/JCM.42.1.408-411.2004
Impact of DNA polymerases and their buffer systems on quantitative real-time PCR
Abstract
An investigation of the influence of five DNA polymerase-buffer systems on real-time PCR showed that the choice of both DNA polymerase and the buffer system affected the amplification efficiency as well as the detection window. The analytical repeatability of the data for different systems changed clearly, leading us to conclude that basing quantitative measurements on single-data-set standard curves can lead to significant errors.
Figures

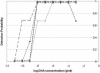
References
-
- Hein, J., U. Schellenberg, G. Bein, and H. Hackstein. 2001. Quantification of murine IFN-gamma mRNA and protein expression: impact of real-time kinetic RT-PCR using SYBR green I dye. Scand. J. Immunol. 54:285-291. - PubMed
-
- Ibekwe, A. M., and C. M. Grieve. 2003. Detection and quantification of Escherichia coli O157:H7 in environmental samples by real-time PCR. J. Appl. Microbiol. 94:421-431. - PubMed
-
- Klein, D., P. Janda, R. Steinborn, M. Muller, B. Salmons, and W. H. Gunzburg. 1999. Proviral load determination of different feline immunodeficiency virus isolates using real-time polymerase chain reaction: influence of mismatches on quantification. Electrophoresis 20:291-299. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

