High-resolution molecular structure of a peptide in an amyloid fibril determined by magic angle spinning NMR spectroscopy
- PMID: 14715898
- PMCID: PMC321745
- DOI: 10.1073/pnas.0304849101
High-resolution molecular structure of a peptide in an amyloid fibril determined by magic angle spinning NMR spectroscopy
Abstract
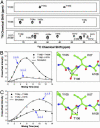
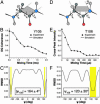
Amyloid fibrils are self-assembled filamentous structures associated with protein deposition conditions including Alzheimer's disease and the transmissible spongiform encephalopathies. Despite the immense medical importance of amyloid fibrils, no atomic-resolution structures are available for these materials, because the intact fibrils are insoluble and do not form diffraction-quality 3D crystals. Here we report the high-resolution structure of a peptide fragment of the amyloidogenic protein transthyretin, TTR(105-115), in its fibrillar form, determined by magic angle spinning NMR spectroscopy. The structure resolves not only the backbone fold but also the precise conformation of the side chains. Nearly complete (13)C and (15)N resonance assignments for TTR(105-115) formed the basis for the extraction of a set of distance and dihedral angle restraints. A total of 76 self-consistent experimental measurements, including 41 restraints on 19 backbone dihedral angles and 35 (13)C-(15)N distances between 3 and 6 A were obtained from 2D and 3D NMR spectra recorded on three fibril samples uniformly (13)C, (15)N-labeled in consecutive stretches of four amino acids and used to calculate an ensemble of peptide structures. Our results indicate that TTR(105-115) adopts an extended beta-strand conformation in the amyloid fibrils such that both the main- and side-chain torsion angles are close to their optimal values. Moreover, the structure of this peptide in the fibrillar form has a degree of long-range order that is generally associated only with crystalline materials. These findings provide an explanation of the unusual stability and characteristic properties of this form of polypeptide assembly.
Figures

References
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

