Bunyamwera bunyavirus RNA synthesis requires cooperation of 3'- and 5'-terminal sequences
- PMID: 14722268
- PMCID: PMC321414
- DOI: 10.1128/jvi.78.3.1129-1138.2004
Bunyamwera bunyavirus RNA synthesis requires cooperation of 3'- and 5'-terminal sequences
Abstract
Bunyamwera virus (BUNV) is the prototype of both the Orthobunyavirus genus and the Bunyaviridae family of segmented negative-sense RNA viruses. The tripartite BUNV genome consists of small (S), medium (M), and large (L) segments that are each transcribed to yield a single mRNA and are replicated to generate an antigenome that acts as a template for synthesis of further genomic strands. As for all negative-sense RNA viruses, the 3'- and 5'-terminal nontranslated regions (NTRs) of the BUNV S, M, and L segments exhibit nucleotide complementarity and, except for one conserved U-G pairing, this complementarity extends for 15, 18, and 19 nucleotides, respectively. We investigated whether the complementarity of 3' and 5' NTRs reflected a functional requirement for terminal cooperation to promote BUNV RNA synthesis or, alternatively, was a consequence of genomic and antigenomic NTRs having similar functions requiring sequence conservation. We show that cooperation between 3'- and 5'-NTR sequences is required for BUNV RNA synthesis, and our results suggest that this cooperation is due to nucleotide complementarity allowing 3' and 5' NTRs to associate through base-pairing interactions. To examine the importance of complementarity in promoting BUNV RNA synthesis, we utilized a competitive replication assay able to examine the replication ability of all possible combinations of interacting nucleotides within a defined region of BUNV 3' and 5' NTRs. We show here that maximal RNA replication was signaled when sequences exhibiting perfect complementarity within 3' and 5' NTRs were selected.
Figures
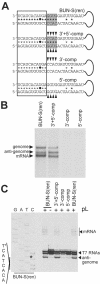

References
-
- Barr, J. N., R. M. Elliott, E. F. Dunn, and G. W. Wertz. 2003. Segment-specific terminal sequences of Bunyamwera bunyavirus regulate genome replication. Virology 311:326-338. - PubMed
-
- Bouloy, M., N. Pardigon, P. Vialat, S. Gerbaud, and M. Girard. 1990. Characterization of the 5′ and 3′ ends of viral messenger RNAs isolated from BHK21 cells infected with Germiston virus (Bunyavirus). Virology 175:50-58. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

