The conserved RNA recognition motif 3 of U2 snRNA auxiliary factor (U2AF 65) is essential in vivo but dispensable for activity in vitro
- PMID: 14730023
- PMCID: PMC1370536
- DOI: 10.1261/rna.5153204
The conserved RNA recognition motif 3 of U2 snRNA auxiliary factor (U2AF 65) is essential in vivo but dispensable for activity in vitro
Abstract
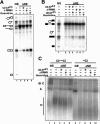
The general splicing factor U2AF(65) recognizes the polypyrimidine tract (Py tract) that precedes 3' splice sites and has three RNA recognition motifs (RRMs). The C-terminal RRM (RRM3), which is highly conserved, has been proposed to contribute to Py-tract binding and establish protein-protein contacts with splicing factors mBBP/SF1 and SAP155. Unexpectedly, we find that the human RRM3 domain is dispensable for U2AF(65) activity in vitro. However, it has an essential function in Schizosaccharomyces pombe distinct from binding to the Py tract or to mBBP/SF1 and SAP155. First, deletion of RRM3 from the human protein has no effect on Py-tract binding. Second, RRM123 and RRM12 select similar sequences from a random pool of RNA. Third, deletion of RRM3 has no effect on the splicing activity of U2AF(65) in vitro. However, deletion of the RRM3 domain of S. pombe U2AF(59) abolishes U2AF function in vivo. In addition, certain amino acid substitutions on the four-stranded beta-sheet surface of RRM3 compromise U2AF function in vivo without affecting binding to mBBP/SF1 or SAP155 in vitro. We propose that RRM3 has an unrecognized function that is possibly relevant for the splicing of only a subset of cellular introns. We discuss the implications of these observations on previous models of U2AF function.
Figures







References
-
- Abovich, N., Liao, X.C., and Rosbash, M. 1994. The yeast MUD2 protein: An interaction with PRP11 defines a bridge between commitment complexes and U2 snRNP addition. Genes & Dev. 8: 843–854. - PubMed
-
- Bashaw, G.J. and Baker, B.S. 1997. The regulation of the Drosophila msl-2 gene reveals a function for Sex-lethal in translational control. Cell 89: 789–798. - PubMed
-
- Bennett, M., Michaud, S., Kingston, J., and Reed, R. 1992. Protein components specifically associated with prespliceosome and spliceosome complexes. Genes & Dev. 6: 1986–2000. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
