O-glycosylation as a sorting determinant for cell surface delivery in yeast
- PMID: 14742720
- PMCID: PMC379253
- DOI: 10.1091/mbc.e03-07-0511
O-glycosylation as a sorting determinant for cell surface delivery in yeast
Abstract
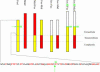
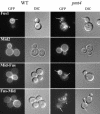
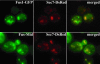
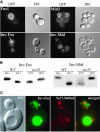
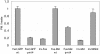
Little is known about the mechanisms that determine localization of proteins to the plasma membrane in Saccharomyces cerevisiae. The length of the transmembrane domains and association of proteins with lipid rafts have been proposed to play a role in sorting to the cell surface. Here, we report that Fus1p, an O-glycosylated integral membrane protein involved in cell fusion during yeast mating, requires O-glycosylation for cell surface delivery. In cells lacking PMT4, encoding a mannosyltransferase involved in the initial step of O-glycosylation, Fus1p was not glycosylated and accumulated in late Golgi structures. A chimeric protein lacking O-glycosylation motif was missorted to the vacuole and accumulated in late Golgi in wild-type cells. Exocytosis of this protein could be restored by addition of a 33-amino acid portion of an O-glycosylated sequence from Fus1p. Our data suggest that O-glycosylation functions as a sorting determinant for cell surface delivery of Fus1p.
Figures

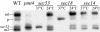


References
-
- Alfalah, M., Jacob, R., Preuss, U., Zimmer, K.P., Naim, H., and Naim, H.Y. (1999). O-Linked glycans mediate apical sorting of human intestinal sucraseisomaltase through association with lipid rafts. Curr. Biol. 9, 593-596. - PubMed
-
- Babczinski, P., and Tanner, W. (1973). Involvement of dolicholmonophosphate in the formation of specific mannosyl-linkages in yeast glycoproteins. Biochem. Biophys. Res. Commun. 54, 1119-1124. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

