Recognition of DNA substrates by T4 bacteriophage polynucleotide kinase
- PMID: 14754987
- PMCID: PMC373337
- DOI: 10.1093/nar/gkh212
Recognition of DNA substrates by T4 bacteriophage polynucleotide kinase
Abstract
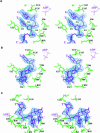
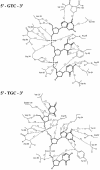
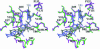
T4 phage polynucleotide kinase (PNK) displays 5'-hydroxyl kinase, 3'-phosphatase and 2',3'-cyclic phosphodiesterase activities. The enzyme phosphorylates the 5' hydroxyl termini of a wide variety of nucleic acid substrates, a behavior studied here through the determination of a series of crystal structures with single-stranded (ss)DNA oligonucleotide substrates of various lengths and sequences. In these structures, the 5' ribose hydroxyl is buried in the kinase active site in proper alignment for phosphoryl transfer. Depending on the ssDNA length, the first two or three nucleotide bases are well ordered. Numerous contacts are made both to the phosphoribosyl backbone and to the ordered bases. The position, side chain contacts and internucleotide stacking interactions of the ordered bases are strikingly different for a 5'-GT DNA end than for a 5'-TG end. The base preferences displayed at those positions by PNK are attributable to differences in the enzyme binding interactions and in the DNA conformation for each unique substrate molecule.
Figures

Similar articles
-
Structure of a tRNA repair enzyme and molecular biology workhorse: T4 polynucleotide kinase.Structure. 2002 Sep;10(9):1249-60. doi: 10.1016/s0969-2126(02)00835-3. Structure. 2002. PMID: 12220496
-
Structure and mechanism of T4 polynucleotide kinase: an RNA repair enzyme.EMBO J. 2002 Jul 15;21(14):3873-80. doi: 10.1093/emboj/cdf397. EMBO J. 2002. PMID: 12110598 Free PMC article.
-
A nanoplatform based on metal-organic frameworks and coupled exonuclease reaction for the fluorimetric determination of T4 polynucleotide kinase activity and inhibition.Mikrochim Acta. 2020 Mar 23;187(4):243. doi: 10.1007/s00604-020-4194-y. Mikrochim Acta. 2020. PMID: 32206934
-
Construction of linker-scanning mutations by oligonucleotide ligation.Methods Mol Biol. 1996;57:279-85. doi: 10.1385/0-89603-332-5:279. Methods Mol Biol. 1996. PMID: 8850014 Review. No abstract available.
-
Polynucleotide kinase as a potential target for enhancing cytotoxicity by ionizing radiation and topoisomerase I inhibitors.Anticancer Agents Med Chem. 2008 May;8(4):358-67. doi: 10.2174/187152008784220311. Anticancer Agents Med Chem. 2008. PMID: 18473721 Free PMC article. Review.
Cited by
-
Investigation into the annotation of protocol sequencing steps in the sequence read archive.Gigascience. 2015 May 9;4:23. doi: 10.1186/s13742-015-0064-7. eCollection 2015. Gigascience. 2015. PMID: 25960871 Free PMC article.
-
Reconstitution and structure of a bacterial Pnkp1-Rnl-Hen1 RNA repair complex.Nat Commun. 2015 Apr 17;6:6876. doi: 10.1038/ncomms7876. Nat Commun. 2015. PMID: 25882814 Free PMC article.
-
Structure-function analysis of the 3' phosphatase component of T4 polynucleotide kinase/phosphatase.Virology. 2007 Sep 15;366(1):126-36. doi: 10.1016/j.virol.2007.03.059. Epub 2007 May 9. Virology. 2007. PMID: 17493655 Free PMC article.
-
Structural and biochemical analysis of the phosphate donor specificity of the polynucleotide kinase component of the bacterial pnkp•hen1 RNA repair system.Biochemistry. 2013 Jul 9;52(27):4734-43. doi: 10.1021/bi400412x. Epub 2013 Jun 26. Biochemistry. 2013. PMID: 23721485 Free PMC article.
-
Structures of bacterial polynucleotide kinase in a Michaelis complex with GTP•Mg2+ and 5'-OH oligonucleotide and a product complex with GDP•Mg2+ and 5'-PO4 oligonucleotide reveal a mechanism of general acid-base catalysis and the determinants of phosphoacceptor recognition.Nucleic Acids Res. 2014 Jan;42(2):1152-61. doi: 10.1093/nar/gkt936. Epub 2013 Oct 22. Nucleic Acids Res. 2014. PMID: 24150947 Free PMC article.
References
-
- Novogrodsky A. and Hurwitz,J. (1966) The enzymatic phosphorylation of ribonucleic acid and deoxyribonucleic acid. I. Phosphorylation at 5′-hydroxyl termini. J. Biol. Chem., 241, 2923–2932. - PubMed
-
- Novogrodsky A., Tal,M., Traub,A. and Hurwitz,J. (1966) The enzymatic phosphorylation of ribonucleic acid and deoxyribonucleic acid. II. Further properties of the 5′-hydroxyl polynucleotide kinase. J. Biol. Chem., 241, 2933–2943. - PubMed
-
- Berkner K.L. and Folk,W.R. (1980) Polynucleotide kinase exchange as an assay for class II restriction endonucleases. Methods Enzymol., 65, 28–36. - PubMed
-
- Chaconas G. and van de Sande,J.H. (1980) 5′-32P labeling of RNA and DNA restriction fragments. Methods Enzymol., 65, 75–85. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

