In silico identification and experimental validation of PmrAB targets in Salmonella typhimurium by regulatory motif detection
- PMID: 14759259
- PMCID: PMC395753
- DOI: 10.1186/gb-2004-5-2-r9
In silico identification and experimental validation of PmrAB targets in Salmonella typhimurium by regulatory motif detection
Abstract
Background: The PmrAB (BasSR) two-component regulatory system is required for Salmonella typhimurium virulence. PmrAB-controlled modifications of the lipopolysaccharide (LPS) layer confer resistance to cationic antibiotic polypeptides, which may allow bacteria to survive within macrophages. The PmrAB system also confers resistance to Fe3+-mediated killing. New targets of the system have recently been discovered that seem not to have a role in the well-described functions of PmrAB, suggesting that the PmrAB-dependent regulon might contain additional, unidentified targets.
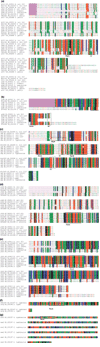
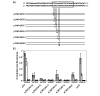
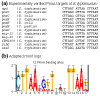
Results: We performed an in silico analysis of possible targets of the PmrAB system. Using a motif model of the PmrA binding site in DNA, genome-wide screening was carried out to detect PmrAB target genes. To increase confidence in the predictions, all putative targets were subjected to a cross-species comparison (phylogenetic footprinting) using a Gibbs sampling-based motif-detection procedure. As well as the known targets, we detected additional targets with unknown functions. Four of these were experimentally validated (yibD, aroQ, mig-13 and sseJ). Site-directed mutagenesis of the PmrA-binding site (PmrA box) in yibD revealed specific sequence requirements.
Conclusions: We demonstrated the efficiency of our procedure by recovering most of the known PmrAB-dependent targets and by identifying unknown targets that we were able to validate experimentally. We also pinpointed directions for further research that could help elucidate the S. typhimurium virulence pathway.
Figures

Similar articles
-
PhoP-PhoQ activates transcription of pmrAB, encoding a two-component regulatory system involved in Salmonella typhimurium antimicrobial peptide resistance.J Bacteriol. 1996 Dec;178(23):6857-64. doi: 10.1128/jb.178.23.6857-6864.1996. J Bacteriol. 1996. PMID: 8955307 Free PMC article.
-
Cationic antimicrobial peptides serve as activation signals for the Salmonella Typhimurium PhoPQ and PmrAB regulons in vitro and in vivo.Front Cell Infect Microbiol. 2012 Jul 27;2:102. doi: 10.3389/fcimb.2012.00102. eCollection 2012. Front Cell Infect Microbiol. 2012. PMID: 22919691 Free PMC article.
-
Genome-wide analysis of the PreA/PreB (QseB/QseC) regulon of Salmonella enterica serovar Typhimurium.BMC Microbiol. 2009 Feb 23;9:42. doi: 10.1186/1471-2180-9-42. BMC Microbiol. 2009. PMID: 19236707 Free PMC article.
-
The Salmonella PmrAB regulon: lipopolysaccharide modifications, antimicrobial peptide resistance and more.Trends Microbiol. 2008 Jun;16(6):284-90. doi: 10.1016/j.tim.2008.03.007. Epub 2008 May 6. Trends Microbiol. 2008. PMID: 18467098 Review.
-
Enhancer-dependent transcription in Salmonella enterica Typhimurium: new members of the sigmaN regulon inferred from protein sequence homology and predicted promoter sites.J Mol Microbiol Biotechnol. 2002 Jul;4(4):367-74. J Mol Microbiol Biotechnol. 2002. PMID: 12125817 Review.
Cited by
-
Adrenaline modulates the global transcriptional profile of Salmonella revealing a role in the antimicrobial peptide and oxidative stress resistance responses.BMC Genomics. 2008 Oct 6;9:458. doi: 10.1186/1471-2164-9-458. BMC Genomics. 2008. PMID: 18837991 Free PMC article.
-
The effect of orthology and coregulation on detecting regulatory motifs.PLoS One. 2010 Feb 3;5(2):e8938. doi: 10.1371/journal.pone.0008938. PLoS One. 2010. PMID: 20140085 Free PMC article.
-
Comparative genomic analysis uncovers 3 novel loci encoding type six secretion systems differentially distributed in Salmonella serotypes.BMC Genomics. 2009 Aug 4;10:354. doi: 10.1186/1471-2164-10-354. BMC Genomics. 2009. PMID: 19653904 Free PMC article.
-
A PmrB-Regulated Deacetylase Required for Lipid A Modification and Polymyxin Resistance in Acinetobacter baumannii.Antimicrob Agents Chemother. 2015 Dec;59(12):7911-4. doi: 10.1128/AAC.00515-15. Epub 2015 Oct 12. Antimicrob Agents Chemother. 2015. PMID: 26459891 Free PMC article.
-
Effects of Regulatory Network Organization and Environment on PmrD Connector Activity and Polymyxin Resistance in Klebsiella pneumoniae and Escherichia coli.Antimicrob Agents Chemother. 2021 Feb 17;65(3):e00889-20. doi: 10.1128/AAC.00889-20. Print 2021 Feb 17. Antimicrob Agents Chemother. 2021. PMID: 33361295 Free PMC article.
References
-
- Gunn JS, Ryan SS, Van Velkinburgh JC, Ernst RK, Miller SI. Genetic and functional analysis of a PmrA-PmrB-regulated locus necessary for lipopolysaccharide modification, antimicrobial peptide resistance, and oral virulence of Salmonella enterica serovar typhimurium. Infect Immun. 2000;68:6139–6146. doi: 10.1128/IAI.68.11.6139-6146.2000. - DOI - PMC - PubMed
-
- Wosten MM, Kox LF, Chamnongpol S, Soncini FC, Groisman EA. A signal transduction system that responds to extracellular iron. Cell. 2000;103:113–125. - PubMed
-
- Garcia Vescovi E, Soncini FC, Groisman EA. Mg2+ as an extracellular signal: environmental regulation of Salmonella virulence. Cell. 1996;84:165–174. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

