Structural basis of HP1/PXVXL motif peptide interactions and HP1 localisation to heterochromatin
- PMID: 14765118
- PMCID: PMC1271814
- DOI: 10.1038/sj.emboj.7600088
Structural basis of HP1/PXVXL motif peptide interactions and HP1 localisation to heterochromatin
Abstract
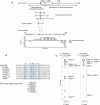
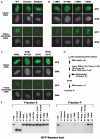
HP1 family proteins are adaptor molecules, containing two related chromo domains that are required for chromatin packaging and gene silencing. Here we present the structure of the chromo shadow domain from mouse HP1beta bound to a peptide containing a consensus PXVXL motif found in many HP1 binding partners. The shadow domain exhibits a novel mode of peptide recognition, where the peptide binds across the dimer interface, sandwiched in a beta-sheet between strands from each monomer. The structure allows us to predict which other shadow domains bind similar PXVXL motif-containing peptides and provides a framework for predicting the sequence specificity of the others. We show that targeting of HP1beta to heterochromatin requires shadow domain interactions with PXVXL-containing proteins in addition to chromo domain recognition of Lys-9-methylated histone H3. Interestingly, it also appears to require the simultaneous recognition of two Lys-9-methylated histone H3 molecules. This finding implies a further complexity to the histone code for regulation of chromatin structure and suggests how binding of HP1 family proteins may lead to its condensation.
Figures


References
-
- Bannister AJ, Zegerman P, Partridge JF, Miska EA, Thomas JO, Allshire RC, Kouzarides T (2001) Selective recognition of methylated lysine 9 on histone H3 by the HP1 chromo domain. Nature 410: 120–124 - PubMed
-
- Brown KE, Baxter J, Graf D, Merkenschlager M, Fisher AG (1999) Dynamic repositioning of genes in the nucleus of lymphocytes preparing for cell division. Mol Cell 3: 207–217 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

