Sequence saturation mutagenesis (SeSaM): a novel method for directed evolution
- PMID: 14872057
- PMCID: PMC373423
- DOI: 10.1093/nar/gnh028
Sequence saturation mutagenesis (SeSaM): a novel method for directed evolution
Abstract
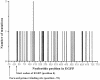
Sequence saturation mutagenesis (SeSaM) is a conceptually novel and practically simple method that truly randomizes a target sequence at every single nucleotide position. A SeSaM experiment can be accomplished within 2-3 days and comprises four steps: generating a pool of DNA fragments with random length, 'tailing' the DNA fragments with universal base using terminal transferase at 3'-termini, elongating DNA fragments in a PCR to the full-length genes using a single-stranded template and replacing the universal bases by standard nucleotides. Random mutations are created at universal sites due to the promiscuous base-pairing property of universal bases. Using enhanced green fluorescence protein as the model system and deoxyinosine as the universal base, we proved by sequencing 100 genes the concept of the SeSaM method and achieved a random distribution of mutations with the mutational bias expected for deoxyinosine.
Figures







References
-
- Brakmann S. (2001) Discovery of superior enzymes by directed molecular evolution. Chem. Biochem., 2, 865–871. - PubMed
-
- Cadwell R.C. and Joyce,G.F. (1992) Randomization of genes by PCR mutagenesis. PCR Methods Appl., 2, 28–33. - PubMed
-
- Cadwell R.C. and Joyce,G.F. (1994) Mutagenic PCR. PCR Methods Appl., 3, 136–140. - PubMed
-
- Lin-Goerke J.L., Robbins,D.J. and Burczak,J.D. (1997) PCR-based random mutagenesis using manganese and reduced dNTP concentration. Biotechniques, 23, 409–412. - PubMed
-
- Zaccolo M., Williams,D.M., Brown,D.M. and Gherardi,E. (1996) An approach to random mutagenesis of DNA using mixtures of triphosphate derivatives of nucleoside analogues. J. Mol. Biol., 255, 589–603. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

