The DNA-A component of a plant geminivirus (Indian mung bean yellow mosaic virus) replicates in budding yeast cells
- PMID: 14963136
- PMCID: PMC369238
- DOI: 10.1128/jvi.78.5.2405-2413.2004
The DNA-A component of a plant geminivirus (Indian mung bean yellow mosaic virus) replicates in budding yeast cells
Abstract
Understanding the biochemistry of DNA replication of the plant DNA viruses is important for the development of antiviral strategies. Since DNA replication is little studied in plants, a genetically tractable, easily culturable, eukaryotic model system is required to pursue such studies in a facile manner. Here we report the development of a yeast model system that supports DNA replication of a chosen geminivirus strain, Indian mung bean yellow mosaic virus. The replication of plasmid DNA in the model system relies specifically on the virus-derived elements and factors. Usage of this model system revealed the role of at least one hitherto unknown viral factor for viral DNA replication. The episomal characteristic of single-strandedness of replicated plasmid DNA was shown, and the expression of viral genes was also confirmed. This model system is expected to shed light on the machinery and mechanism involved in geminiviral DNA replication in plants.
Figures
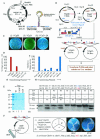



Similar articles
-
A nanovirus-like DNA component associated with yellow vein disease of Ageratum conyzoides: evidence for interfamilial recombination between plant DNA viruses.Virology. 1999 Nov 10;264(1):142-52. doi: 10.1006/viro.1999.9948. Virology. 1999. PMID: 10544139
-
Agrobacterium tumefaciens supports DNA replication of diverse geminivirus types.FEBS Lett. 2002 Apr 10;516(1-3):179-82. doi: 10.1016/s0014-5793(02)02539-5. FEBS Lett. 2002. PMID: 11959128
-
A versatile transreplication-based system to identify cellular proteins involved in geminivirus replication.J Virol. 2006 Apr;80(7):3624-33. doi: 10.1128/JVI.80.7.3624-3633.2006. J Virol. 2006. PMID: 16537630 Free PMC article.
-
Insights into the functional characteristics of geminivirus rolling-circle replication initiator protein and its interaction with host factors affecting viral DNA replication.Arch Virol. 2015 Feb;160(2):375-87. doi: 10.1007/s00705-014-2297-7. Epub 2014 Dec 2. Arch Virol. 2015. PMID: 25449306 Review.
-
Engineering resistance to geminiviruses--review and perspectives.Plant Biotechnol J. 2007 Mar;5(2):207-20. doi: 10.1111/j.1467-7652.2006.00217.x. Plant Biotechnol J. 2007. PMID: 17309676 Review.
Cited by
-
Saccharomyces cerevisiae: a versatile eukaryotic system in virology.Microb Cell Fact. 2007 Oct 10;6:32. doi: 10.1186/1475-2859-6-32. Microb Cell Fact. 2007. PMID: 17927824 Free PMC article.
-
Yeast for virus research.Microb Cell. 2017 Sep 18;4(10):311-330. doi: 10.15698/mic2017.10.592. Microb Cell. 2017. PMID: 29082230 Free PMC article. Review.
-
Diversity and recombination analysis of Cotton leaf curl Multan virus: a highly emerging begomovirus in northern India.BMC Genomics. 2019 Apr 6;20(1):274. doi: 10.1186/s12864-019-5640-2. BMC Genomics. 2019. PMID: 30954067 Free PMC article.
-
Arabidopsis thaliana NAC083 protein interacts with Mungbean yellow mosaic India virus (MYMIV) Rep protein.Virus Genes. 2014 Jun;48(3):486-93. doi: 10.1007/s11262-013-1028-6. Epub 2014 Jan 19. Virus Genes. 2014. PMID: 24442717
-
Formation of AAV single stranded DNA genome from a circular plasmid in Saccharomyces cerevisiae.PLoS One. 2011;6(8):e23474. doi: 10.1371/journal.pone.0023474. Epub 2011 Aug 10. PLoS One. 2011. PMID: 21853137 Free PMC article.
References
-
- Agatep, R., R. D. Kirkpatrick, D. L. Parchaliuk, R. A. Woods, and R. D. Gietz. 1998. Transformation of Saccharomyces cerevisiae by lithium acetate/single stranded DNA/polyethylene glycol (LiAc/ss-DNA/PEG) protocol. Technical Tips Online [Online.] http://tto.trends.com.
-
- Ausubel, F. M., R. Brent, R. E. Kingston, D. D. Moore, J. G. Seidman, J. A. Smith, and K. Struhl. 1998. Current protocols in molecular biology, vol. 2. John Wiley & Sons, Inc., London, United Kingdom.
-
- Castillo, A. G., D. Collinet, S. Deret, A. Kashoggi, and E. R. Bejarrano. 2003. Dual interaction of plant PCNA with geminivirus replication accessory protein (Ren) and viral replication protein (Rep). Virology 312:381-394. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

