DNA methylation may restrict but does not determine differential gene expression at the Sgy/Tead2 locus during mouse development
- PMID: 14966277
- PMCID: PMC350557
- DOI: 10.1128/MCB.24.5.1968-1982.2004
DNA methylation may restrict but does not determine differential gene expression at the Sgy/Tead2 locus during mouse development
Abstract
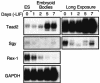
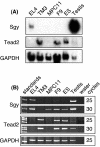
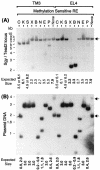
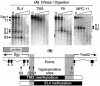
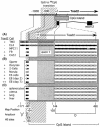
Soggy (Sgy) and Tead2, two closely linked genes with CpG islands, were coordinately expressed in mouse preimplantation embryos and embryonic stem (ES) cells but were differentially expressed in differentiated cells. Analysis of established cell lines revealed that Sgy gene expression could be fully repressed by methylation of the Sgy promoter and that DNA methylation acted synergistically with chromatin deacetylation. Differential gene expression correlated with differential DNA methylation, resulting in sharp transitions from methylated to unmethylated DNA at the open promoter in both normal cells and tissues, as well as in established cell lines. However, neither promoter was methylated in normal cells and tissues even when its transcripts were undetectable. Moreover, the Sgy promoter remained unmethylated as Sgy expression was repressed during ES cell differentiation. Therefore, DNA methylation was not the primary determinant of Sgy/Tead2 expression. Nevertheless, Sgy expression was consistently restricted to basal levels whenever downstream regulatory sequences were methylated, suggesting that DNA methylation restricts but does not regulate differential gene expression during mouse development.
Figures





References
-
- Adachi, N., and M. R. Lieber. 2002. Bidirectional gene organization: a common architectural feature of the human genome. Cell 109:807-809. - PubMed
-
- Baylin, S. B., M. Esteller, M. R. Rountree, K. E. Bachman, K. Schuebel, and J. G. Herman. 2001. Aberrant patterns of DNA methylation, chromatin formation and gene expression in cancer. Hum. Mol. Genet. 10:687-692. - PubMed
-
- Brady, G., and N. N. Iscove. 1993. Construction of cDNA libraries from single cells. Methods Enzymol. 225:611-623. - PubMed
-
- Cameron, E. E., K. E. Bachman, S. Myohanen, J. G. Herman, and S. B. Baylin. 1999. Synergy of demethylation and histone deacetylase inhibition in the re-expression of genes silenced in cancer. Nat. Genet. 21:103-107. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
