Promyelocytic leukemia nuclear bodies associate with transcriptionally active genomic regions
- PMID: 14970191
- PMCID: PMC2171989
- DOI: 10.1083/jcb.200305142
Promyelocytic leukemia nuclear bodies associate with transcriptionally active genomic regions
Abstract
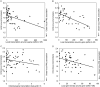
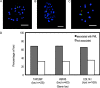
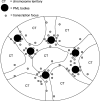
The promyelocytic leukemia (PML) protein is aggregated into nuclear bodies that are associated with diverse nuclear processes. Here, we report that the distance between a locus and its nearest PML body correlates with the transcriptional activity and gene density around the locus. Genes on the active X chromosome are more significantly associated with PML bodies than their silenced homologues on the inactive X chromosome. We also found that a histone-encoding gene cluster, which is transcribed only in S-phase, is more strongly associated with PML bodies in S-phase than in G0/G1 phase of the cell cycle. However, visualization of specific RNA transcripts for several genes showed that PML bodies were not themselves sites of transcription for these genes. Furthermore, knock-down of PML bodies by RNA interference did not preferentially change the expression of genes closely associated with PML bodies. We propose that PML bodies form in nuclear compartments of high transcriptional activity, but they do not directly regulate transcription of genes in these compartments.
Copyright The Rockefeller University Press
Figures





Similar articles
-
PML bodies: a meeting place for genomic loci?J Cell Sci. 2005 Mar 1;118(Pt 5):847-54. doi: 10.1242/jcs.01700. J Cell Sci. 2005. PMID: 15731002
-
Cell cycle-dependent association of PML bodies with sites of active transcription in nuclei of mammalian cells.J Struct Biol. 2002 Oct-Dec;140(1-3):167-79. doi: 10.1016/s1047-8477(02)00571-3. J Struct Biol. 2002. PMID: 12490165
-
PML bodies associate specifically with the MHC gene cluster in interphase nuclei.J Cell Sci. 2001 Oct;114(Pt 20):3705-16. doi: 10.1242/jcs.114.20.3705. J Cell Sci. 2001. PMID: 11707522
-
Pondering the puzzle of PML (promyelocytic leukemia) nuclear bodies: can we fit the pieces together using an RNA regulon?Biochim Biophys Acta. 2008 Nov;1783(11):2145-54. doi: 10.1016/j.bbamcr.2008.06.005. Epub 2008 Jun 18. Biochim Biophys Acta. 2008. PMID: 18616965 Free PMC article. Review.
-
Targeting promyelocytic leukemia protein: a means to regulating PML nuclear bodies.Int J Biol Sci. 2009 May 22;5(4):366-76. doi: 10.7150/ijbs.5.366. Int J Biol Sci. 2009. PMID: 19471587 Free PMC article. Review.
Cited by
-
SUMO-1-dependent allosteric regulation of thymine DNA glycosylase alters subnuclear localization and CBP/p300 recruitment.Mol Cell Biol. 2007 Jan;27(1):229-43. doi: 10.1128/MCB.00323-06. Epub 2006 Oct 23. Mol Cell Biol. 2007. PMID: 17060459 Free PMC article.
-
TIN2 deficiency leads to ALT-associated phenotypes and differentiation defects in embryonic stem cells.Stem Cell Reports. 2022 May 10;17(5):1183-1197. doi: 10.1016/j.stemcr.2022.03.005. Epub 2022 Apr 7. Stem Cell Reports. 2022. PMID: 35395177 Free PMC article.
-
A manually curated network of the PML nuclear body interactome reveals an important role for PML-NBs in SUMOylation dynamics.Int J Biol Sci. 2010 Jan 12;6(1):51-67. doi: 10.7150/ijbs.6.51. Int J Biol Sci. 2010. PMID: 20087442 Free PMC article. Review.
-
The dynamic organization of gene-regulatory machinery in nuclear microenvironments.EMBO Rep. 2005 Feb;6(2):128-33. doi: 10.1038/sj.embor.7400337. EMBO Rep. 2005. PMID: 15689940 Free PMC article. Review.
-
The Arc of cognition: Signaling cascades regulating Arc and implications for cognitive function and disease.Semin Cell Dev Biol. 2018 May;77:63-72. doi: 10.1016/j.semcdb.2017.09.023. Semin Cell Dev Biol. 2018. PMID: 29559111 Free PMC article. Review.

