TeiR, a LuxR-type transcription factor required for testosterone degradation in Comamonas testosteroni
- PMID: 14973025
- PMCID: PMC344414
- DOI: 10.1128/JB.186.5.1430-1437.2004
TeiR, a LuxR-type transcription factor required for testosterone degradation in Comamonas testosteroni
Abstract
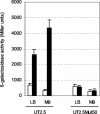
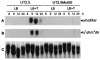
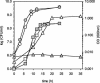
We have identified a new steroid-inducible gene (designated teiR [testosterone-inducible regulator]) in Comamonas testosteroni that is required for testosterone degradation. Nucleotide sequence analysis of teiR predicts a 391-amino-acid protein which shows homology between residues 327 and 380 (C-terminal domain) to the LuxR helix-turn-helix DNA binding domain and between residues 192 and 227 to the PAS sensor domain. This domain distribution resembles that described for TraR, a specific transcriptional regulator involved in quorum sensing in Agrobacterium tumefaciens. Analysis of the gene expression indicated that teiR is tightly controlled at the transcriptional level by the presence of testosterone in the culture medium. A teiR-disrupted mutant strain was completely unable to use testosterone as the sole carbon and energy source. In addition, the expression of several steroid-inducible genes was abolished in this mutant. Northern blot assays revealed that teiR is required for full expression of sip48-beta-HSD gene mRNA (encoding a steroid-inducible protein of 48 kDa and 3beta-17beta-hydroxysteroid dehydrogenase) and also of other steroid degradation genes, including those encoding 3alpha-hydroxysteroid dehydrogenase, Delta(5)-3-ketoisomerase, 3-oxo-steroid Delta(1)-dehydrogenase, and 3-oxo-steroid Delta(4)-(5alpha)-dehydrogenase enzymes. Moreover, when teiR was provided to the teiR-disrupted strain in trans, the transcription level of these genes was restored. These results indicate that TeiR positively regulates the transcription of genes involved in the initial enzymatic steps of steroid degradation in C. testosteroni.
Figures


Similar articles
-
Testosterone-inducible regulator is a kinase that drives steroid sensing and metabolism in Comamonas testosteroni.J Biol Chem. 2008 Jun 20;283(25):17380-90. doi: 10.1074/jbc.M710166200. Epub 2008 Apr 17. J Biol Chem. 2008. PMID: 18424443
-
Regulation of testosterone degradation in Comamonas testosteroni.J Steroid Biochem Mol Biol. 2008 Nov;112(1-3):145-50. doi: 10.1016/j.jsbmb.2008.09.011. Epub 2008 Sep 21. J Steroid Biochem Mol Biol. 2008. PMID: 18852046
-
Identification of a novel steroid inducible gene associated with the beta hsd locus of Comamonas testosteroni.J Steroid Biochem Mol Biol. 2004 Jan;88(1):91-100. doi: 10.1016/j.jsbmb.2003.10.010. J Steroid Biochem Mol Biol. 2004. PMID: 15026087
-
Comprehensive review of steroid metabolism in Comamonas testosteroni TA441 with insights from other aerobic steroid-degrading bacteria.Adv Appl Microbiol. 2025;131:1-20. doi: 10.1016/bs.aambs.2025.06.001. Epub 2025 Aug 7. Adv Appl Microbiol. 2025. PMID: 40818839 Review.
-
Role and regulation of bacterial LuxR-like regulators.J Cell Biochem. 2011 Oct;112(10):2694-702. doi: 10.1002/jcb.23219. J Cell Biochem. 2011. PMID: 21678467 Review.
Cited by
-
Steroids as Environmental Compounds Recalcitrant to Degradation: Genetic Mechanisms of Bacterial Biodegradation Pathways.Genes (Basel). 2019 Jul 6;10(7):512. doi: 10.3390/genes10070512. Genes (Basel). 2019. PMID: 31284586 Free PMC article. Review.
-
Identification of 9α-hydroxy-17-oxo-1,2,3,4,10,19-hexanorandrostan-5-oic acid in steroid degradation by Comamonas testosteroni TA441 and its conversion to the corresponding 6-en-5-oyl coenzyme A (CoA) involving open reading frame 28 (ORF28)- and ORF30-encoded acyl-CoA dehydrogenases.J Bacteriol. 2014 Oct;196(20):3598-608. doi: 10.1128/JB.01878-14. Epub 2014 Aug 4. J Bacteriol. 2014. PMID: 25092028 Free PMC article.
-
Permanent draft genome sequence of Comamonas testosteroni KF-1.Stand Genomic Sci. 2013 May 30;8(2):239-54. doi: 10.4056/sigs.3847890. eCollection 2013. Stand Genomic Sci. 2013. PMID: 23991256 Free PMC article.
-
Identification of "missing links" in C- and D-ring cleavage of steroids by Comamonas testosteroni TA441.Appl Environ Microbiol. 2023 Oct 31;89(10):e0105023. doi: 10.1128/aem.01050-23. Epub 2023 Oct 10. Appl Environ Microbiol. 2023. PMID: 37815342 Free PMC article.
-
Identification of the EdcR Estrogen-Dependent Repressor in Caenibius tardaugens NBRC 16725: Construction of a Cellular Estradiol Biosensor.Genes (Basel). 2021 Nov 23;12(12):1846. doi: 10.3390/genes12121846. Genes (Basel). 2021. PMID: 34946795 Free PMC article.
References
-
- Abalain, J., S. Di Stefano, M. L. Abalain-Colloc, and H. H. Floch. 1995. Cloning, sequencing and expression of Pseudomonas testosteroni gene encoding 3α-hydroxysteroid dehydrogenase. J. Steroid Biochem. Mol. Biol. 55:233-238. - PubMed
-
- Abalain, J. H., S. Di Stéfano, Y. Amet, E. Quemeneur, M. L. Abalain-Colloc, and H. H. Floch. 1993. Cloning, DNA sequencing and expression of (3-17)-β hydroxysteroid dehydrogenase from Pseudomonas testosteroni. J. Steroid Biochem. Mol. Biol. 44:133-139. - PubMed
-
- Ahmad, D., R. Masse, and M. Sylvestre. 1990. Cloning and expression of genes involved in 4-chlorobiphenyl transformation by Pseudomonas testosteroni, homology to polychlorobiphenyl-degrading genes in other bacteria. Gene 86:53-61. - PubMed
-
- Altschul, S. F., W. Gish, E. Miller, E. W. Myers, and D. J. Lipman. 1990. Basic local alignment search tool. J. Mol. Biol. 215:403-410. - PubMed
-
- Arai, H., S. Akahira, T. Ohishi, and T. Kudo. 1999. Adaptation of Comamonas testosteroni TA441 to utilization of phenol by spontaneous mutation of the gene for a trans acting factor. Mol. Microbiol. 33:1132-1140. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

