A single amino acid of APOBEC3G controls its species-specific interaction with virion infectivity factor (Vif)
- PMID: 14978281
- PMCID: PMC374346
- DOI: 10.1073/pnas.0307132101
A single amino acid of APOBEC3G controls its species-specific interaction with virion infectivity factor (Vif)
Abstract
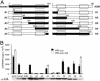
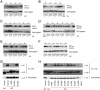
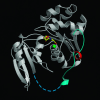
The virion infectivity factor (Vif) accessory protein of HIV-1 forms a complex with the cellular cytidine deaminase APOBEC3G (apolipoprotein B mRNA-editing enzyme, catalytic polypeptide-like 3G) to block its antiviral activity. The antiviral property of APOBEC3G is conserved in several mammalian species, but the ability of Vif to block this activity is species-specific. HIV-1 Vif blocks human APOBEC3G but does not block the mouse or African green monkey (AGM) enzyme. Conversely, SIV(AGM) Vif blocks the antiviral activity of AGM but not human APOBEC3G. We demonstrate that the species specificity is caused by a single amino acid difference in APOBEC3G. Replacement of Asp-128 in human APOBEC3G with the Lys-128 of AGM APOBEC3G caused the enzyme to switch its interaction, becoming sensitive to SIV(AGM) Vif and resistant to HIV-1 Vif. Conversely, the reciprocal Lys to Asp switch in AGM APOBEC3G reversed its specificity for Vif. The reversal of biological activity was accompanied by the corresponding switch in the species specificity with which the enzyme physically associated with Vif and was excluded from virions. The charge of the amino acid at position 128 was a critical determinant of species specificity. Based on the crystal structure of the distantly related Escherichia coli cytidine deaminase, we propose that this amino acid is positioned on a solvent-exposed loop of APOBEC3G on the same face of the protein as the catalytic site.
Figures

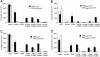
Comment in
-
Controlling lentiviruses: single amino acid changes can determine specificity.Proc Natl Acad Sci U S A. 2004 Mar 16;101(11):3725-6. doi: 10.1073/pnas.0400929101. Epub 2004 Mar 9. Proc Natl Acad Sci U S A. 2004. PMID: 15010528 Free PMC article. No abstract available.
Similar articles
-
A single amino acid difference in the host APOBEC3G protein controls the primate species specificity of HIV type 1 virion infectivity factor.Proc Natl Acad Sci U S A. 2004 Mar 16;101(11):3770-4. doi: 10.1073/pnas.0307713101. Epub 2004 Mar 3. Proc Natl Acad Sci U S A. 2004. PMID: 14999100 Free PMC article.
-
A single amino acid substitution in human APOBEC3G antiretroviral enzyme confers resistance to HIV-1 virion infectivity factor-induced depletion.Proc Natl Acad Sci U S A. 2004 Apr 13;101(15):5652-7. doi: 10.1073/pnas.0400830101. Epub 2004 Mar 30. Proc Natl Acad Sci U S A. 2004. PMID: 15054139 Free PMC article.
-
A single amino acid determinant governs the species-specific sensitivity of APOBEC3G to Vif action.J Biol Chem. 2004 Apr 9;279(15):14481-3. doi: 10.1074/jbc.C400060200. Epub 2004 Feb 13. J Biol Chem. 2004. PMID: 14966139
-
New insights into the role of Vif in HIV-1 replication.AIDS Rev. 2004 Jan-Mar;6(1):34-9. AIDS Rev. 2004. PMID: 15168739 Review.
-
Cellular interactions of virion infectivity factor (Vif) as potential therapeutic targets: APOBEC3G and more?Curr Drug Targets. 2006 Dec;7(12):1583-93. doi: 10.2174/138945006779025356. Curr Drug Targets. 2006. PMID: 17168833 Review.
Cited by
-
Random mutagenesis MAPPIT analysis identifies binding sites for Vif and Gag in both cytidine deaminase domains of Apobec3G.PLoS One. 2012;7(9):e44143. doi: 10.1371/journal.pone.0044143. Epub 2012 Sep 10. PLoS One. 2012. PMID: 22970171 Free PMC article.
-
Antiviral Mechanism and Biochemical Basis of the Human APOBEC3 Family.Front Microbiol. 2012 Jul 9;3:250. doi: 10.3389/fmicb.2012.00250. eCollection 2012. Front Microbiol. 2012. PMID: 22787460 Free PMC article.
-
Epitranscriptomic addition of m6A regulates HIV-1 RNA stability and alternative splicing.Genes Dev. 2021 Jul 1;35(13-14):992-1004. doi: 10.1101/gad.348508.121. Epub 2021 Jun 17. Genes Dev. 2021. PMID: 34140354 Free PMC article.
-
APOBEC3G and HIV-1: strike and counterstrike.Curr Infect Dis Rep. 2006 Jun;8(4):317-23. doi: 10.1007/s11908-006-0077-6. Curr Infect Dis Rep. 2006. PMID: 16822376
-
Structural Basis for a Species-Specific Determinant of an SIV Vif Protein toward Hominid APOBEC3G Antagonism.Cell Host Microbe. 2019 Dec 11;26(6):739-747.e4. doi: 10.1016/j.chom.2019.10.014. Cell Host Microbe. 2019. PMID: 31830442 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

