Analysis and prediction of affinity of TAP binding peptides using cascade SVM
- PMID: 14978300
- PMCID: PMC2286721
- DOI: 10.1110/ps.03373104
Analysis and prediction of affinity of TAP binding peptides using cascade SVM
Abstract
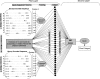
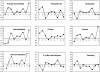
The generation of cytotoxic T lymphocyte (CTL) epitopes from an antigenic sequence involves number of intracellular processes, including production of peptide fragments by proteasome and transport of peptides to endoplasmic reticulum through transporter associated with antigen processing (TAP). In this study, 409 peptides that bind to human TAP transporter with varying affinity were analyzed to explore the selectivity and specificity of TAP transporter. The abundance of each amino acid from P1 to P9 positions in high-, intermediate-, and low-affinity TAP binders were examined. The rules for predicting TAP binding regions in an antigenic sequence were derived from the above analysis. The quantitative matrix was generated on the basis of contribution of each position and residue in binding affinity. The correlation of r = 0.65 was obtained between experimentally determined and predicted binding affinity by using a quantitative matrix. Further a support vector machine (SVM)-based method has been developed to model the TAP binding affinity of peptides. The correlation (r = 0.80) was obtained between the predicted and experimental measured values by using sequence-based SVM. The reliability of prediction was further improved by cascade SVM that uses features of amino acids along with sequence. An extremely good correlation (r = 0.88) was obtained between measured and predicted values, when the cascade SVM-based method was evaluated through jackknife testing. A Web service, TAPPred (http://www.imtech.res.in/raghava/tappred/ or http://bioinformatics.uams.edu/mirror/tappred/), has been developed based on this approach.
Figures



Similar articles
-
TAPPred prediction of TAP-binding peptides in antigens.Methods Mol Biol. 2007;409:381-6. doi: 10.1007/978-1-60327-118-9_28. Methods Mol Biol. 2007. PMID: 18450016
-
Selection and binding of peptides to human transporters associated with antigen processing and rat cim-a and -b.J Immunol. 1996 Jul 1;157(1):213-20. J Immunol. 1996. PMID: 8683118
-
Targeting tumour cells with defects in the MHC Class I antigen processing pathway with CD8+ T cells specific for hydrophobic TAP- and Tapasin-independent peptides: the requirement for directed access into the ER.Cancer Immunol Immunother. 2007 Aug;56(8):1143-52. doi: 10.1007/s00262-006-0263-2. Epub 2006 Dec 2. Cancer Immunol Immunother. 2007. PMID: 17143611 Free PMC article.
-
A comprehensive review and performance evaluation of bioinformatics tools for HLA class I peptide-binding prediction.Brief Bioinform. 2020 Jul 15;21(4):1119-1135. doi: 10.1093/bib/bbz051. Brief Bioinform. 2020. PMID: 31204427 Free PMC article. Review.
-
Function of the transport complex TAP in cellular immune recognition.Biochim Biophys Acta. 1999 Dec 6;1461(2):405-19. doi: 10.1016/s0005-2736(99)00171-6. Biochim Biophys Acta. 1999. PMID: 10581370 Review.
Cited by
-
Modeling the structure of bound peptide ligands to major histocompatibility complex.Protein Sci. 2004 Sep;13(9):2523-32. doi: 10.1110/ps.04631204. Protein Sci. 2004. PMID: 15322290 Free PMC article.
-
The Importance of Being Presented: Target Validation by Immunopeptidomics for Epitope-Specific Immunotherapies.Front Immunol. 2022 Apr 6;13:883989. doi: 10.3389/fimmu.2022.883989. eCollection 2022. Front Immunol. 2022. PMID: 35464395 Free PMC article. Review.
-
Systems Immunology: Learning the Rules of the Immune System.Annu Rev Immunol. 2018 Apr 26;36:813-842. doi: 10.1146/annurev-immunol-042617-053035. Annu Rev Immunol. 2018. PMID: 29677477 Free PMC article. Review.
-
A computational assay to design an epitope-based Peptide vaccine against saint louis encephalitis virus.Bioinform Biol Insights. 2013 Nov 24;7:347-55. doi: 10.4137/BBI.S13402. eCollection 2013. Bioinform Biol Insights. 2013. PMID: 24324329 Free PMC article.
-
In silico analysis of STX2a-PE15-P4A8 chimeric protein as a novel immunotoxin for cancer therapy.In Silico Pharmacol. 2021 Feb 10;9(1):19. doi: 10.1007/s40203-021-00079-w. eCollection 2021. In Silico Pharmacol. 2021. PMID: 33643767 Free PMC article.
References
-
- Abele, R. and Tampe, R. 1999. Function of the transport complex TAP in cellular immune recognition. Biochim. Biophys. Acta. 1461 405–419. - PubMed
-
- Androlewicz, M.J. and Cresswell, P. 1994. Human transporters associated with antigen processing possess a promiscuous peptide-binding site. Immunity 1 7–14. - PubMed
-
- Bhasin, M., Singh, H., and Raghava, G.P.S. 2003. MHCBN: A comprehensive database of MHC binding and non-binding peptides. Bioinformatics 19 666–667. - PubMed
-
- Bhaskaran, R. and Ponnuswamy, P.K. 1988. Positional flexibilities of amino acid residues in globular proteins. Int. J. Pept. Protein Res. 32 241–255. - PubMed
-
- Blythe, M.J., Doytchinova, I.A., and Flower, D.R. 2002. JenPep: A database of quantitative functional peptide data for immunology. Bioinformatics 18 434–439. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

