Comparison of BRCT domains of BRCA1 and 53BP1: a biophysical analysis
- PMID: 14978302
- PMCID: PMC2286730
- DOI: 10.1110/ps.03461404
Comparison of BRCT domains of BRCA1 and 53BP1: a biophysical analysis
Abstract
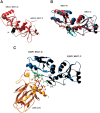
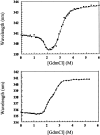
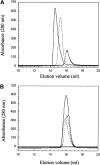
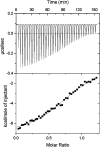
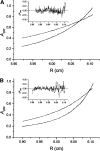
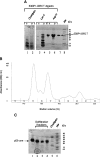
53BP1 interacts with the DNA-binding core domain of the tumor suppressor p53 and enhances p53-mediated transcriptional activation. The p53-binding region of 53BP1 maps to the C-terminal BRCT domains, which are homologous to those found in the breast cancer protein BRCA1 and in other proteins involved in DNA repair. Here we compare the thermodynamic behavior of the BRCT domains of 53BP1 and BRCA1 and examine their ability to interact with the p53 core domain. The free energies of unfolding are of similar magnitude, although slightly higher for 53BP1-BRCT, and both populate an aggregation-prone partly folded intermediate. Interaction studies performed in vitro by analytical size-exclusion chromatography, analytical ultracentrifugation, and isothermal titration calorimetry reveal that 53BP1-BRCT interacts with p53 with a K(d) in the low micromolar range. Despite their homology with 53BP1-BRCT domains, the BRCT domains of BRCA1 did not bind p53 with any detectable affinity. In summary, although other studies have indicated that the BRCT domains of both BRCA1 and 53BP1 interact with p53 core domain, the quantitative biophysical measurements performed here indicate that only 53BP1 can bind. Although both proteins may be involved in the same DNA repair pathways, our study indicates that a direct role in p53 function is unique to 53BP1.
Figures
References
-
- Chai, Y.L., Cui, J., Shao, N., Shyam, E., Reddy, P., and Rao, V.N. 1999. The second BRCT domain of BRCA1 proteins interacts with p53 and stimulates transcription from the p21WAF1/CIP1 promoter. Oncogene 18 263–268. - PubMed
-
- DiTullio Jr., R.A., Mochan, T.A., Venere, M., Bartkova, J., Sehested, M., Bartek, J., and Halazonetis, T.D. 2002. 53BP1 functions in an ATM-dependent checkpoint pathway that is constitutively activated in human cancer. Nat. Cell Biol. 4 998–1002. - PubMed
-
- Ekblad, C.M., Wilkinson, H.R., Schymkowitz, J.W., Rousseau, F., Freund, S.M., and Itzhaki, L.S. 2002. Characterisation of the BRCT domains of the breast cancer susceptibility gene product BRCA1. J. Mol. Biol. 320 431–442. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

