From conversion to aggregation: protofibril formation of the prion protein
- PMID: 14983003
- PMCID: PMC356944
- DOI: 10.1073/pnas.0307178101
From conversion to aggregation: protofibril formation of the prion protein
Abstract
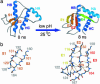
The ability to diagnose and treat prion diseases is limited by our current understanding of the conversion process of the protein from healthy to harmful isoform. Whereas the monomeric, benign species is well characterized, the misfolded conformations responsible for infectivity and neurodegeneration remain elusive. There is mounting evidence that fibrillization intermediates, or protofibrils, but not mature fibrils or plaques, are the pathogenic species in amyloid diseases. Here, we use molecular dynamics to simulate the conversion of the prion protein. Molecular dynamics simulation produces a scrapie prion protein-like conformation enriched in beta-structure that is in good agreement with available experimental data. The converted conformation was then used to model a protofibril by means of the docking of hydrophobic patches of the template structure to form hydrogen-bonded sheets spanning adjacent subunits. The resulting protofibril model provides a non-branching aggregate with a 3(1) axis of symmetry that is in good agreement with a wide variety of experimental data; importantly, it was derived from realistic simulation of the conversion process.
Figures



Similar articles
-
Formation of soluble oligomers and amyloid fibrils with physical properties of the scrapie isoform of the prion protein from the C-terminal domain of recombinant murine prion protein mPrP-(121-231).J Biol Chem. 2006 Sep 8;281(36):26121-8. doi: 10.1074/jbc.M605367200. Epub 2006 Jul 13. J Biol Chem. 2006. PMID: 16844683
-
Molecular modelling indicates that the pathological conformations of prion proteins might be beta-helical.Biochem J. 1999 Oct 15;343 Pt 2(Pt 2):453-60. Biochem J. 1999. PMID: 10510313 Free PMC article.
-
Nucleation-dependent conformational conversion of the Y145Stop variant of human prion protein: structural clues for prion propagation.Proc Natl Acad Sci U S A. 2003 Oct 14;100(21):12069-74. doi: 10.1073/pnas.2033281100. Epub 2003 Sep 30. Proc Natl Acad Sci U S A. 2003. PMID: 14519851 Free PMC article.
-
Conformational diseases and structure-toxicity relationships: lessons from prion-derived peptides.Curr Protein Pept Sci. 2007 Feb;8(1):83-90. doi: 10.2174/138920307779941505. Curr Protein Pept Sci. 2007. PMID: 17305562 Review.
-
Conformational conversion of prion protein in prion diseases.Acta Biochim Biophys Sin (Shanghai). 2013 Jun;45(6):465-76. doi: 10.1093/abbs/gmt027. Epub 2013 Apr 11. Acta Biochim Biophys Sin (Shanghai). 2013. PMID: 23580591 Review.
Cited by
-
Generating Bona Fide Mammalian Prions with Internal Deletions.J Virol. 2016 Jul 11;90(15):6963-6975. doi: 10.1128/JVI.00555-16. Print 2016 Aug 1. J Virol. 2016. PMID: 27226369 Free PMC article.
-
The role of α-sheet structure in amyloidogenesis: characterization and implications.Open Biol. 2022 Nov;12(11):220261. doi: 10.1098/rsob.220261. Epub 2022 Nov 23. Open Biol. 2022. PMID: 36416010 Free PMC article. Review.
-
Segments in the Amyloid Core that Distinguish Hamster from Mouse Prion Fibrils.Neurochem Res. 2019 Jun;44(6):1399-1409. doi: 10.1007/s11064-018-02709-w. Epub 2019 Jan 2. Neurochem Res. 2019. PMID: 30603982
-
Theoretical model of prion propagation: a misfolded protein induces misfolding.Proc Natl Acad Sci U S A. 2005 May 31;102(22):7835-40. doi: 10.1073/pnas.0409389102. Epub 2005 May 23. Proc Natl Acad Sci U S A. 2005. PMID: 15911770 Free PMC article.
-
Misfolding pathways of the prion protein probed by molecular dynamics simulations.Biophys J. 2005 Feb;88(2):1334-43. doi: 10.1529/biophysj.104.049882. Epub 2004 Nov 19. Biophys J. 2005. PMID: 15556981 Free PMC article.
References
-
- Prusiner, S. B. (1991) Science 252, 1515–1522. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

