Temperature-dependent conformational transitions and hydrogen-bond dynamics of the elastin-like octapeptide GVG(VPGVG): a molecular-dynamics study
- PMID: 14990469
- PMCID: PMC1303977
- DOI: 10.1016/S0006-3495(04)74210-1
Temperature-dependent conformational transitions and hydrogen-bond dynamics of the elastin-like octapeptide GVG(VPGVG): a molecular-dynamics study
Abstract
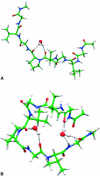
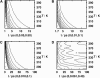
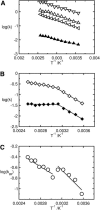
A joint experimental/theoretical investigation of the elastin-like octapeptide GVG(VPGVG) was carried out. In this article a comprehensive molecular-dynamics study of the temperature-dependent folding and unfolding of the octapeptide is presented. The current study, as well as its experimental counterpart (see companion article in this issue) find that this peptide undergoes an inverse temperature transition (ITT), leading to a folding at approximately 40-60 degrees C. In addition, an unfolding transition is identified at unusually high temperatures approaching the normal boiling point of water. Due to the small size of the system, two broad temperature regimes are found: the ITT regime at approximately 10-60 degrees C and the unfolding regime at approximately T > 60 degrees C, where the peptide has a maximum probability of being folded at T approximately 60 degrees C. A detailed molecular picture involving a thermodynamic order parameter, or reaction coordinate, for this process is presented along with a time-correlation function analysis of the hydrogen-bond dynamics within the peptide as well as between the peptide and solvating water molecules. Correlation with experimental evidence and ramifications on the properties of elastin are discussed.
Figures






Similar articles
-
Characterization of the temperature- and pressure-induced inverse and reentrant transition of the minimum elastin-like polypeptide GVG(VPGVG) by DSC, PPC, CD, and FT-IR spectroscopy.Biophys J. 2004 Mar;86(3):1385-92. doi: 10.1016/S0006-3495(04)74209-5. Biophys J. 2004. PMID: 14990468 Free PMC article.
-
Inverse temperature transition of a biomimetic elastin model: reactive flux analysis of folding/unfolding and its coupling to solvent dielectric relaxation.J Phys Chem B. 2006 Mar 2;110(8):3576-87. doi: 10.1021/jp054805a. J Phys Chem B. 2006. PMID: 16494413
-
Folding and unfolding of an elastinlike oligopeptide: "inverse temperature transition," reentrance, and hydrogen-bond dynamics.Phys Rev Lett. 2004 Apr 9;92(14):148101. doi: 10.1103/PhysRevLett.92.148101. Epub 2004 Apr 7. Phys Rev Lett. 2004. PMID: 15089575
-
[A turning point in the knowledge of the structure-function-activity relations of elastin].J Soc Biol. 2001;195(2):181-93. J Soc Biol. 2001. PMID: 11727705 Review. French.
-
Molecular basis for the extensibility of elastin.J Muscle Res Cell Motil. 2002;23(5-6):561-73. doi: 10.1023/a:1023474909980. J Muscle Res Cell Motil. 2002. PMID: 12785105 Review.
Cited by
-
A water-explicit lattice model of heat-, cold-, and pressure-induced protein unfolding.Biophys J. 2007 Dec 15;93(12):4116-27. doi: 10.1529/biophysj.107.108530. Epub 2007 Aug 31. Biophys J. 2007. PMID: 17766342 Free PMC article.
-
Improving Internal Peptide Dynamics in the Coarse-Grained MARTINI Model: Toward Large-Scale Simulations of Amyloid- and Elastin-like Peptides.J Chem Theory Comput. 2012 May 8;8(5):1774-1785. doi: 10.1021/ct200876v. Epub 2012 Mar 26. J Chem Theory Comput. 2012. PMID: 22582033 Free PMC article.
-
Coarse-grained model of tropoelastin self-assembly into nascent fibrils.Mater Today Bio. 2019 Jun 18;3:100016. doi: 10.1016/j.mtbio.2019.100016. eCollection 2019 Jun. Mater Today Bio. 2019. PMID: 32159149 Free PMC article.
-
Computational smart polymer design based on elastin protein mutability.Biomaterials. 2017 May;127:49-60. doi: 10.1016/j.biomaterials.2017.01.041. Epub 2017 Jan 31. Biomaterials. 2017. PMID: 28279921 Free PMC article.
-
Solid-State NMR characterization of autofluorescent fibrils formed by the elastin-derived peptide GVGVAGVG.Biomacromolecules. 2011 May 9;12(5):1546-55. doi: 10.1021/bm101486s. Epub 2011 Apr 1. Biomacromolecules. 2011. PMID: 21456595 Free PMC article.
References
-
- Aalten, D. V., B. D. Groot, J. Findlay, H. Berendsen, and A. Amadei. 1997. A comparison of techniques for calculating protein essential dynamics. J. Comp. Chem. 18:169–181.
-
- Aaron, B., and J. Gosline. 1980. Optical properties of single elastin fibres indicate random protein conformation. Nature. 287:865–867. - PubMed
-
- Amadei, A., A. Linssen, and H. Berendsen. 1993. Essential dynamics of proteins. Proteins Struct. Funct. Gen. 17:412–425. - PubMed
-
- Andriciolaei, I., and M. Karplus. 2001. On the calculation of entropy from covariance matrices of the atomic fluctuations. J. Chem. Phys. 115:6289–6292.
-
- Arad, M. G. O. 1990. Depsipeptide analogues of elastin repeating sequences: conformational analysis. Biopolymers. 29:1651–1668. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

