Control of yeast filamentous-form growth by modules in an integrated molecular network
- PMID: 14993204
- PMCID: PMC353223
- DOI: 10.1101/gr.2020604
Control of yeast filamentous-form growth by modules in an integrated molecular network
Abstract
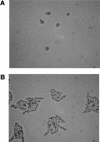
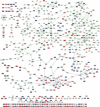
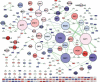
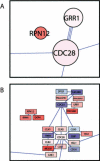
On solid growth media with limiting nitrogen source, diploid budding-yeast cells differentiate from the yeast form to a filamentous, adhesive, and invasive form. Genomic profiles of mRNA levels in Saccharomyces cerevisiae yeast-form and filamentous-form cells were compared. Disparate data types, including genes implicated by expression change, filamentation genes known previously through a phenotype, protein-protein interaction data, and protein-metabolite interaction data were integrated as the nodes and edges of a filamentation-network graph. Application of a network-clustering method revealed 47 clusters in the data. The correspondence of the clusters to modules is supported by significant coordinated expression change among cluster co-member genes, and the quantitative identification of collective functions controlling cell properties. The modular abstraction of the filamentation network enables the association of filamentous-form cell properties with the activation or repression of specific biological processes, and suggests hypotheses. A module-derived hypothesis was tested. It was found that the 26S proteasome regulates filamentous-form growth.
Figures

Similar articles
-
Identification of translational regulation target genes during filamentous growth in Saccharomyces cerevisiae: regulatory role of Caf20 and Dhh1.Eukaryot Cell. 2006 Dec;5(12):2120-7. doi: 10.1128/EC.00121-06. Epub 2006 Oct 13. Eukaryot Cell. 2006. PMID: 17041186 Free PMC article.
-
[Selective elimination of meiosis-specific mRNA operates during the mitotic cell cycle].Tanpakushitsu Kakusan Koso. 2007 Jan;52(1):1-10. Tanpakushitsu Kakusan Koso. 2007. PMID: 17228834 Review. Japanese. No abstract available.
-
Systematic identification of pathways that couple cell growth and division in yeast.Science. 2002 Jul 19;297(5580):395-400. doi: 10.1126/science.1070850. Epub 2002 Jun 27. Science. 2002. PMID: 12089449
-
[Rpn4p is a positive and negative transcriptional regulator of the ubiquitin-proteasome system].Mol Biol (Mosk). 2008 May-Jun;42(3):518-25. Mol Biol (Mosk). 2008. PMID: 18702311 Russian.
-
Yeast Gal4: a transcriptional paradigm revisited.EMBO Rep. 2006 May;7(5):496-9. doi: 10.1038/sj.embor.7400679. EMBO Rep. 2006. PMID: 16670683 Free PMC article. Review.
Cited by
-
Control of signaling in a MAP-kinase pathway by an RNA-binding protein.PLoS One. 2007 Feb 28;2(2):e249. doi: 10.1371/journal.pone.0000249. PLoS One. 2007. PMID: 17327913 Free PMC article.
-
Prediction of phenotype and gene expression for combinations of mutations.Mol Syst Biol. 2007;3:96. doi: 10.1038/msb4100137. Epub 2007 Mar 27. Mol Syst Biol. 2007. PMID: 17389876 Free PMC article.
-
Multiple signals converge on a differentiation MAPK pathway.PLoS Genet. 2010 Mar 19;6(3):e1000883. doi: 10.1371/journal.pgen.1000883. PLoS Genet. 2010. PMID: 20333241 Free PMC article.
-
The KUPNetViz: a biological network viewer for multiple -omics datasets in kidney diseases.BMC Bioinformatics. 2013 Jul 24;14:235. doi: 10.1186/1471-2105-14-235. BMC Bioinformatics. 2013. PMID: 23883183 Free PMC article.
-
The promoter of filamentation (POF1) protein from Saccharomyces cerevisiae is an ATPase involved in the protein quality control process.BMC Microbiol. 2011 Dec 28;11:268. doi: 10.1186/1471-2180-11-268. BMC Microbiol. 2011. PMID: 22204397 Free PMC article.
References
-
- Bajorek, M., Finley, D., and Glickman, M.H. 2003. Proteasome disassembly and downregulation is correlated with viability during stationary phase. Curr. Biol. 13: 1140–1144. - PubMed
-
- Costanzo, M.C., Crawford, M.E., Hirschman, J.E., Kranz, J.E., Olsen, P., Robertson, L.S., Skrzypek, M.S., Braun, B.R., Hopkins, K.L., Kondu, P., et al. 2001. YPD, PombePD and WormPD: Model organism volumes of the BioKnowledge library, an integrated resource for protein information. Nucleic Acids Res. 29: 75–79. - PMC - PubMed
WEB SITE REFERENCES
-
- http://www.incyte.com/control/tools/proteome; Incyte Yeast Protein Database.
-
- http://labs.systemsbiology.net/galitski; Galitski Lab home page.
-
- http://www.ncbi.nlm.nih.gov/geo/; Gene Expression Omnibus database.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
