Solution structure of dengue virus capsid protein reveals another fold
- PMID: 14993605
- PMCID: PMC373476
- DOI: 10.1073/pnas.0305892101
Solution structure of dengue virus capsid protein reveals another fold
Abstract
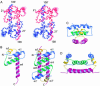
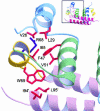
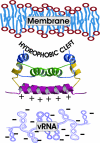
Dengue virus is responsible for approximately 50-100 million infections, resulting in nearly 24,000 deaths annually. The capsid (C) protein of dengue virus is essential for specific encapsidation of the RNA genome, but little structural information on the C protein is available. We report the solution structure of the 200-residue homodimer of dengue 2 C protein. The structure provides, to our knowledge, the first 3D picture of a flavivirus C protein and identifies a fold that includes a large dimerization surface contributed by two pairs of helices, one of which has characteristics of a coiled-coil. NMR structure determination involved a secondary structure sorting approach to facilitate assignment of the intersubunit nuclear Overhauser effect interactions. The dimer of dengue C protein has an unusually high net charge, and the structure reveals an asymmetric distribution of basic residues over the surface of the protein. Nearly half of the basic residues lie along one face of the dimer. In contrast, the conserved hydrophobic region forms an extensive apolar surface at a dimer interface on the opposite side of the molecule. We propose a model for the interaction of dengue C protein with RNA and the viral membrane that is based on the asymmetric charge distribution of the protein and is consistent with previously reported results.
Figures



References
-
- Gubler, D. J. (2002) Trends Microbiol. 10, 100-103. - PubMed
-
- Lindenbach, B. D. & Rice, C. M. (2001) in Fields Virology, eds. Knipe, D. M. & Howley, P. M. (Lippincott, Williams & Wilkins, Philadelphia), pp. 991-1041.
-
- Rey, F. A., Heinz, F. X., Mandl, C., Kunz, C. & Harrison, S. C. (1995) Nature 375, 291-298. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources

