Using pyrrolo-deoxycytosine to probe RNA/DNA hybrids containing the human immunodeficiency virus type-1 3' polypurine tract
- PMID: 15004241
- PMCID: PMC390295
- DOI: 10.1093/nar/gkh307
Using pyrrolo-deoxycytosine to probe RNA/DNA hybrids containing the human immunodeficiency virus type-1 3' polypurine tract
Abstract
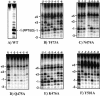
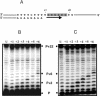
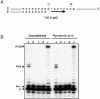
Recent structural analyses indicate that localized regions of abnormal base pairing exist within RNA/DNA hybrids containing the HIV-1 polypurine tract (PPT) and that these distortions may play a role in PPT function. To examine this directly, we have introduced pyrrolo-deoxycytosine (pdC), a fluorescent, environmentally sensitive analog of deoxycytosine (dC), into the DNA strand of PPT-containing hybrids. Steady-state fluorescence analysis of these hybrids reveals that the DNA base 11 nt from the PPT-U3 junction is unpaired even in the absence of reverse transcriptase (RT). Unstable base pairing is also observed within the (rG:dC)6 tract in the downstream portion of the duplex, suggesting that HIV-1 RT may recognize multiple pre-existing distortions during PPT selection. HIV-1 RT hydrolyzes pdC-containing hybrids primarily at the PPT-U3 junction, indicating that the analog does not induce a gross structural deformation of the duplex. However, aberrant cleavage is frequently observed 3 bp from the site of pdC substitution, most likely reflecting a specific interaction between the analog and amino acid residues within the RNase H primer grip. pdC substitution within the template strand of a DNA duplex does not appear to significantly affect RT-catalyzed DNA synthesis. Implications of these findings on the use of pdC to examine nucleic acid structure are discussed.
Figures


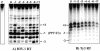

References
-
- Telesnitsky A. and Goff,S.P. (1997) Reverse transcriptase and the generation of retroviral DNA. In Coffin,J.M., Hughes,S.H. and Varmus,H.E. (eds), Retroviruses. Cold Spring Harbor Laboratory Press, Plainview, NY, pp. 121–160. - PubMed
-
- Le Grice S.F. (1993) Human immunodeficiency virus reverse transcriptase. In Skalka,A.M. and Goff,S. (eds), Reverse Transcriptase. Cold Spring Harbor Laboratory Press, Plainview, NY, pp. 163–192.
-
- Hungnes O., Jonsrud,K., Tjotta,E. and Grinde,B. (1993) Sequence comparison and mutational analysis of elements that may be involved in the regulation of DNA synthesis in HIV-1. J. Mol. Evol., 37, 198–203. - PubMed
-
- Wohrl B.M. and Moelling,K. (1990) Interaction of HIV-1 ribonuclease H with polypurine tract containing RNA-DNA hybrids. Biochemistry, 29, 10141–10147. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

