Identifying transcriptional targets
- PMID: 15005803
- PMCID: PMC395755
- DOI: 10.1186/gb-2004-5-3-210
Identifying transcriptional targets
Abstract
Identifying the targets of transcription factors is important for understanding cellular processes. We review how targets have previously been isolated and outline new technologies that are being developed to identify novel direct targets, including chromatin immunoprecipitation combined with microarray screening and bioinformatic approaches.
Figures
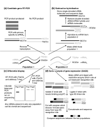
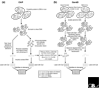
Similar articles
-
Chromatin immunoprecipitation for identifying transcription factor targets in keratinocytes.Methods Mol Biol. 2010;585:159-70. doi: 10.1007/978-1-60761-380-0_12. Methods Mol Biol. 2010. PMID: 19908003
-
Mapping DNA-protein interactions in large genomes by sequence tag analysis of genomic enrichment.Nat Methods. 2005 Jan;2(1):47-53. doi: 10.1038/nmeth726. Epub 2004 Dec 21. Nat Methods. 2005. PMID: 15782160
-
[Finding targets of transcriptional regulators--chromatin immunoprecipitation assay (ChIP)].Postepy Biochem. 2011;57(4):418-24. Postepy Biochem. 2011. PMID: 22568174 Review. Polish.
-
Genome-wide mapping of the binding sites of proteins that interact with DNA.Methods Mol Biol. 2012;881:137-56. doi: 10.1007/978-1-61779-827-6_6. Methods Mol Biol. 2012. PMID: 22639213
-
Visualizing and characterizing in vivo DNA-binding events and direct target genes of plant transcription factors.Methods Mol Biol. 2011;754:293-305. doi: 10.1007/978-1-61779-154-3_17. Methods Mol Biol. 2011. PMID: 21720960 Review.
Cited by
-
Transcription factor HIF1A: downstream targets, associated pathways, polymorphic hypoxia response element (HRE) sites, and initiative for standardization of reporting in scientific literature.Tumour Biol. 2016 Nov;37(11):14851-14861. doi: 10.1007/s13277-016-5331-4. Epub 2016 Sep 19. Tumour Biol. 2016. PMID: 27644243
-
CpG Island microarray probe sequences derived from a physical library are representative of CpG Islands annotated on the human genome.Nucleic Acids Res. 2005 May 23;33(9):2952-61. doi: 10.1093/nar/gki582. Print 2005. Nucleic Acids Res. 2005. PMID: 15911630 Free PMC article.
-
Transcriptomic Analysis of Resistant and Wild-Type Isolates Revealed Fludioxonil as a Candidate for Controlling the Emerging Isoprothiolane Resistant Populations of Magnaporthe oryzae.Front Microbiol. 2022 Apr 8;13:874497. doi: 10.3389/fmicb.2022.874497. eCollection 2022. Front Microbiol. 2022. PMID: 35464942 Free PMC article.
-
Gene regulatory networks in disease and ageing.Nat Rev Nephrol. 2024 Sep;20(9):616-633. doi: 10.1038/s41581-024-00849-7. Epub 2024 Jun 12. Nat Rev Nephrol. 2024. PMID: 38867109 Review.
-
Zebrafish promoter microarrays identify actively transcribed embryonic genes.Genome Biol. 2006;7(8):R71. doi: 10.1186/gb-2006-7-8-R71. Epub 2006 Aug 4. Genome Biol. 2006. PMID: 16889661 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

