Efficient isolation and gene expression profiling of small numbers of neural crest stem cells and developing Schwann cells
- PMID: 15014110
- PMCID: PMC6729482
- DOI: 10.1523/JNEUROSCI.4083-03.2004
Efficient isolation and gene expression profiling of small numbers of neural crest stem cells and developing Schwann cells
Abstract
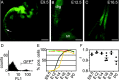
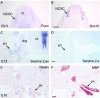
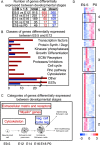
Schwann cells develop from multipotent neural crest stem cells and are important for neuronal survival, maintenance of axonal integrity, and myelination. We used transgenic mice expressing green fluorescent protein in a tissue-specific manner to isolate viable, pure populations of neural crest stem cells and developing Schwann cells, which are not readily accessible by microdissection. Starting with the minute amounts of RNA obtained, a two-round amplification procedure was used to achieve reproducible DNA array hybridizations. We validated our screening procedure by comparisons with the literature and by in situ hybridization. Stage-to-stage comparisons and hierarchical clustering for neural crest and five stages of Schwann cell development suggest a wealth of candidates for genes involved in stem cell regulation and in early Schwann cell development. The combination of methods applied in this study should be generally useful for isolating and profiling other stem cell and difficult to isolate cell populations.
Figures

References
-
- Affymetrix Inc. (2000) Expression analysis: technical manual.
-
- Affymetrix Inc. (2002) GeneChip eukaryotic small sample target labeling assay version II, Technical Note 701265, Rev 3.
-
- Anderson DJ (1997) Cellular and molecular biology of neural crest cell lineage determination. Trends Genet 13: 276-280. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
