A peptide from autoantigen La blocks poliovirus and hepatitis C virus cap-independent translation and reveals a single tyrosine critical for La RNA binding and translation stimulation
- PMID: 15016896
- PMCID: PMC371053
- DOI: 10.1128/jvi.78.7.3763-3776.2004
A peptide from autoantigen La blocks poliovirus and hepatitis C virus cap-independent translation and reveals a single tyrosine critical for La RNA binding and translation stimulation
Abstract
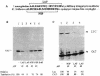
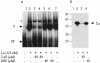
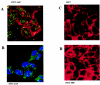
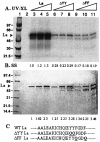
La, a 52-kDa autoantigen in patients with systemic lupus erythematosus, was one of the first cellular proteins identified to interact with viral internal ribosome entry site (IRES) elements and stimulate poliovirus (PV) and hepatitis C virus (HCV) IRES-mediated translation. Previous results from our laboratory have shown that a small, yeast RNA (IRNA) could selectively inhibit PV and HCV IRES-mediated translation by sequestering the La protein. Here we have identified an 18-amino-acid-long sequence from the N-terminal "La motif" which is required for efficient interaction of La with IRNA and viral 5' untranslated region (5'-UTR) elements. A synthetic peptide (called LAP, for La peptide) corresponding to this sequence (amino acids 11 to 28) of La was found to efficiently inhibit viral IRES-mediated translation in vitro. The LAP efficiently enters Huh-7 cells and preferentially inhibits HCV IRES-mediated translation programmed by a bicistronic RNA in vivo. The LAP does not bind RNA directly but appears to block La binding to IRNA and PV 5'-UTR. Competition UV cross-link and translation rescue experiments suggested that LAP inhibits IRES-mediated translation by interacting with proteins rather than RNA. Mutagenesis of LAP demonstrates that single amino acid changes in a highly conserved sequence within LAP are sufficient to eliminate the translation-inhibitory activity of LAP. When one of these mutations (Y23Q) is introduced into full-length La, the mutant protein is severely defective in interacting with the PV IRES element and consequently unable to stimulate IRES-mediated translation. However, the La protein with a mutation of the next tyrosine moiety (Y24Q) could still interact with PV 5'-UTR and stimulate viral IRES-mediated translation significantly. These results underscore the importance of the La N-terminal amino acids in RNA binding and viral RNA translation. The possible role of the LAP sequence in La-RNA binding and stimulation of viral IRES-mediated translation is discussed.
Figures


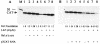

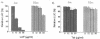

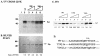
Similar articles
-
Targeting internal ribosome entry site (IRES)-mediated translation to block hepatitis C and other RNA viruses.FEMS Microbiol Lett. 2004 May 15;234(2):189-99. doi: 10.1016/j.femsle.2004.03.045. FEMS Microbiol Lett. 2004. PMID: 15135522 Review.
-
A small yeast RNA blocks hepatitis C virus internal ribosome entry site (HCV IRES)-mediated translation and inhibits replication of a chimeric poliovirus under translational control of the HCV IRES element.J Virol. 1998 Jul;72(7):5638-47. doi: 10.1128/JVI.72.7.5638-5647.1998. J Virol. 1998. PMID: 9621022 Free PMC article.
-
Nucleolin stimulates viral internal ribosome entry site-mediated translation.Virus Res. 2001 Jul;76(1):17-29. doi: 10.1016/s0168-1702(01)00240-4. Virus Res. 2001. PMID: 11376843
-
Human La antigen is required for the hepatitis C virus internal ribosome entry site-mediated translation.J Biol Chem. 2000 Sep 8;275(36):27531-40. doi: 10.1074/jbc.M001487200. J Biol Chem. 2000. PMID: 10856291
-
Inhibition of internal entry site (IRES)-mediated translation by a small yeast RNA: a novel strategy to block hepatitis C virus protein synthesis.Front Biosci. 1998 Dec 1;3:D1241-52. doi: 10.2741/a359. Front Biosci. 1998. PMID: 9835647 Review.
Cited by
-
Versatility of RNA-Binding Proteins in Cancer.Comp Funct Genomics. 2012;2012:178525. doi: 10.1155/2012/178525. Epub 2012 May 14. Comp Funct Genomics. 2012. PMID: 22666083 Free PMC article.
-
A cell-permeable peptide inhibits hepatitis C virus replication by sequestering IRES transacting factors.Virology. 2009 Nov 10;394(1):82-90. doi: 10.1016/j.virol.2009.08.012. Epub 2009 Sep 8. Virology. 2009. PMID: 19740508 Free PMC article.
-
Proteochemometric Method for pIC50 Prediction of Flaviviridae.Biomed Res Int. 2022 Sep 15;2022:7901791. doi: 10.1155/2022/7901791. eCollection 2022. Biomed Res Int. 2022. PMID: 36158882 Free PMC article.
-
Proteome analysis of liver cells expressing a full-length hepatitis C virus (HCV) replicon and biopsy specimens of posttransplantation liver from HCV-infected patients.J Virol. 2005 Jun;79(12):7558-69. doi: 10.1128/JVI.79.12.7558-7569.2005. J Virol. 2005. PMID: 15919910 Free PMC article.
-
Recent advances on the role of host factors during non-poliovirus enteroviral infections.J Biomed Sci. 2019 Jun 19;26(1):47. doi: 10.1186/s12929-019-0540-y. J Biomed Sci. 2019. PMID: 31215493 Free PMC article. Review.
References
-
- Ali, N., and A. Siddiqui. 1997. Molecular mechanisms of translation initiation in eukaryotes. Proc. Natl. Acad. Sci. USA 94:2249-2254. - PubMed
-
- Ali, N., G. J. M. Prujin, D. J. Kenan, J. D. Keene, and A. Siddiqui. 2000. Human La antigen is required for the hepatitis C virus internal ribosome entry site-mediated translation. J. Biol. Chem. 275:27531-27540. - PubMed
-
- Banerjee, R., M. Igo, R. Izumi, U. Datta, and A. Dasgupta. 2000. In vitro replication of RNA viruses, p. 158-164. In A. Cann (ed.), RNA viruses: a practical approach. Oxford University Press, New York, N.Y.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

